1. はじめに
コンピュータ上で化合物の活性や物性を予測することは従来より行われてきたが、特に近年、蓄積データの増大や、Deep Learning技術の発展もあり、AI創薬というキーワードを耳にするようになってきた。
そこで今回、新型コロナウィルス(SARS-CoV-2)の治療薬の予測に関する "Computational Models Identify Several FDA Approved or Experimental Drugs as Putative Agents Against SARS-CoV-2" という論文の内容を自分で試してみることで、AI創薬を**「無料で」**疑似体験してみたので共有したい。
この論文を選んだのは、以下3つの理由による。
- 公共データを利用しており、解析手順、プログラムがgithubで公開されているため、再現しやすい。
- 従来型の解析手法(フィンガープリント、ランダムフォレスト)が用いられており、比較的再現しやすい。
- 新型コロナウィルスの治療薬は、世界中で開発が進められており、興味があった。
2. 今回の作業方針
まず初めに、論文の内容を試すための作業方針を検討した(というか試行錯誤してこうなったというのが正解)
実は本論文における収集データ、解析プログラムは全てgithub( https://github.com/alvesvm/sars-cov-mpro/tree/master/mol-inf-2020 )で公開されており、これを使えば何の苦労もなく論文の内容を再現することができる。
ただ、これをそのままダウンロードし、Jupyter上でリターンキーを連打したとしても、「なるほど」と理解した気になって終わるだけで、何も残らないだろう。
手を動かし、詰まり、調べ、考えることの繰り返しにより、自分の理解不足に気づき、今まで理解できなかったことが、少しずつ理解できるようになると思う。
このため以下の方針で進めることとした。
- 公共データベースからのデータの収集も 自分で調べながら やってみる。
- 収集したデータを用いて、自分で予測プログラムを書いてみる。
- 最終結果を論文と比較し、できれば考察もしてみる。
- 論文の内容の詳細の確認や、詰まった場合のヒントを得る場合、結果の比較のためにgithubを参照する。
次に論文の概要、今回利用する公共データベースについて記載する。
3.参考論文概要
論文では、新型コロナウィルスのウイルス複製に関与するメインプロテアーゼを薬物のターゲット分子としており、これに非常によく似た構造をもつ2003年に流行したSARSコロナウイルスのメインプロテアーゼに対する阻害活性を予測する予測モデルを構築している。
さらに、DrugBankより取得した市販/中止/実験、および治験薬データに対し、構築した予測モデルを適用することで、最終的に41の薬を新型コロナウィルスの候補治療薬として提示している。
なお、学習データとなるSARSコロナウイルスに対する阻害活性情報は、ChEMBLやPDB等の公共データベースより収集している。
ちなみにターゲット分子であるメインプロテアーゼについては、PDBj入門:PDBjの生体高分子学習ポータルサイト 242: コロナウイルスプロテアーゼ(Coronavirus Proteases)で詳しく解説されている。
4. 公共データベースの概要
次に本論文で使われている公共データベースについて簡単にまとめてみる。
ChEMBL (https://www.ebi.ac.uk/chembl/)
ChEMBLは、医薬品および医薬品候補となりうる低分子の生物活性情報について、手動でキュレーションされたものを収載したデータベースである。
欧州分子生物学研究所(EMBL)の欧州バイオインフォマティクス研究所(EBI)によって管理されている。
収載データ件数は次の通りである(ダウンロードしたV27のテーブルのレコード件数より算出)。
| 項目 | 収載件数 |
|---|---|
| 化合物数 | 2,444,828 |
| 生物活性値数 | 16,066,124 |
| アッセイ数 | 1,221,361 |
PDB (https://www.rcsb.org/)
PDB(Protein Data Bank)はタンパク質と核酸の3次元構造データを収載したデータベースである。Worldwide Protein Data Bankにより管理されている。
収載データ件数次のとおりである(2020/9/12現在)。
| 項目 | 収載件数 |
|---|---|
| Structures | 49012 |
| Structures of Human Sequences | 12216 |
| Nucleic Acid Containing Structures | 12216 |
DrugBank (https://www.drugbank.ca/)
DrugBankは、薬物と薬物標的に関する情報を収載したデータベースである。Metabolomics Innovation Center (TMIC)にて管理されている。
最新リリース(バージョン5.1.7、2020-07-02にリリース)の収載件数は以下の通りである。
| 項目 | 収載件数 |
|---|---|
| 薬物 | 13,715 |
| 承認済み小分子薬 | 2,649 |
| 承認済み生物製剤(タンパク質、ペプチド、ワクチン、およびアレルゲン) | 1,405 |
| 栄養補助食品 | 131 |
| 実験 (発見段階)薬物 | 6,417 |
| リンクされたタンパク質シーケンス | 5,234 |
さて、次からいよいよ、論文を内容を試す作業を進める。
5. データ準備
5.1 学習データの準備
ここでは論文に記載されているSARSウィルスのメインプロテアーゼに対する阻害分子の実験データ(ChEMBLE91件、PDB22件)の収集を試みる。
5.1.1 ChEMBLデータの取得
ChEMBLはWEBブラウザでも利用できるが、データをダウンロードし、ローカルのデータベースに登録してSQLから利用することもできる。今回はPostgreSQLをインストールし、PostgreSQL用のダンプをロードして利用することとする。
① ChEMBLデータのダウンロード
ftp://ftp.ebi.ac.uk/pub/databases/chembl/ChEMBLdb/latest/ をクリックするとダウンロード用のファイルの一覧が表示されるので、chembl_27_postgresql.tar.gzをダウンロードし展開する。
② PostgreSQLのインストール
続いてUbuntu18.04にPostgreSQLをインストール 等を参考に、PostgreSQLをインストールする。
③ ChEMBLのロード
PostgreSQLにログインしてデータベースを作成する。
pgdb=# create database chembl_27;
続いてデータを解凍したディレクトリに移動し以下を実行する。
しばらく時間はかかるが、PostgreSQLにデータが取り込まれる。
pg_restore --no-owner -U postgres -d chembl_27 chembl_27_postgresql.dmp
④ ChEMBLのテーブルの理解
次にpgAdmin等からSQLを発行してデータを取得するわけだが、まずはどのテーブルにどんなデータが格納されているのかをftp://ftp.ebi.ac.uk/pub/databases/chembl/ChEMBLdb/latest/schema_documentation.txtをざっと見てみる。またSchema Questions and SQL ExamplesにSQLの例があるのでこれを見て重要そうなテーブルに当たりをつける。あたりをつけたテーブルについて、先のschema_documentation.txtを調べた結果を整理したのが以下の表になる。
| テーブル名 | 説明 |
|---|---|
| compound_structures | 化合物のさまざまな構造表現(Molfile、InChIなど)を格納するテーブル |
| molecule_dictionary | 関連する識別子を持つ化合物/生物学的治療薬の非冗長リスト |
| compound_records | 科学的ドキュメントより抽出された各化合物を表す。 |
| docs | アッセイが抽出されたすべての科学的ドキュメント(論文または特許)を保持する |
| activities | 科学的ドキュメントに記録されたアッセイの結果である活性値またはエンドポイントを格納するテーブル |
| assay | 各ドキュメントで報告されているアッセイのリストを保存する。異なるドキュメントからの同じアッセイは、別のアッセイとして表示される |
| target_dictionary | ChEMBLに含まれる全てのキュレーション済のターゲットの辞書。タンパク質ターゲットと非タンパク質ターゲット(例:生物、組織、細胞株)の両方が含まれる。 |
⑤ データベースにSQLを発行しSARSの阻害データをゲット
ChEMBLでは、全てのデータにCHEMBL IDが割り当てられるため、まずはSARSコロナウィルスのCHEMBL ID検索してみる。このため前項で調べたtarget_dictionaryテーブルに対し以下のSQLを投げてみる。
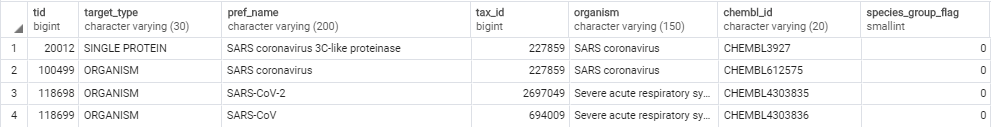
select * from target_dictionary where upper(pref_name) like '%SARS%'
以下の通り4つが得られる。
今回はメインプロテアーゼ(3C-like proteaseとしても知られる)の阻害データを収集したいので、pref_nameより一番上のCHEMBL3927であることが分かる。
ターゲット分子のCHEMBL IDが分かれば、Schema Questions and SQL Examplesの下部の "Retrieve compound activity details for all targets containing a protein of interest" というクエリにより、ターゲット分子に対する活性を持つ分子の一覧が得られるので、このSQLのCHEMBL ID部分を変更すればよい。
SELECT m.chembl_id AS compound_chembl_id,
s.canonical_smiles,
r.compound_key,
d.pubmed_id,
d.doi,
a.description,
act.standard_type,
act.standard_relation,
act.standard_value,
act.standard_units,
act.activity_comment
FROM compound_structures s,
molecule_dictionary m,
compound_records r,
docs d,
activities act,
assays a,
target_dictionary t
WHERE s.molregno = m.molregno
AND m.molregno = r.molregno
AND r.record_id = act.record_id
AND r.doc_id = d.doc_id
AND act.assay_id = a.assay_id
AND a.tid = t.tid
AND t.chembl_id = 'CHEMBL3927'
AND (act.standard_type ='IC50' or act.standard_type ='Ki')
order by compound_chembl_id;
注意点としては今回、阻害データだけほしいので、standard_typeとしてIC50または Kiを指定している。IC50とKiの定義は厳密には異なると思われるが、論文ではともに学習データとして利用しているっぽかったので、ここでは準ずることとする。
結果をpgAdminからcsvにエクスポートしておく。
⑥ データの整理
さて、ここはデータサイエンティストっぽい泥臭い作業となる。
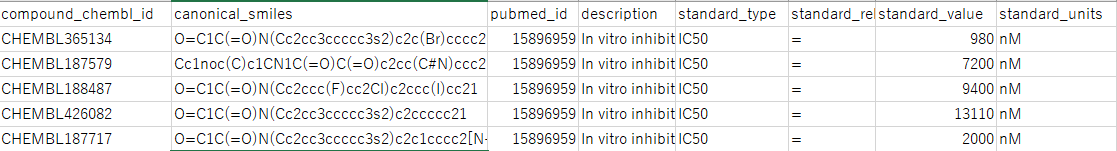
csvには以下のようなデータが得られている。
canonical_smilesは化合物の構造を示すデータであるため、説明変数を生成するために必要となる。
standard_valueはstandard_typeの値(ここでは以降、阻害値と呼ぶことにする)であり、standard_unitsは単位である。
standard_unitsは今回は全部同じnMなので、単位を変換して揃える必要はない。
論文の予測モデル用のデータ作成のため、以下の処理をした。
- csvのstandard_relationに不等号が含まれている場合の対応として、このようなケースでは安全をみた数値として判断することが多いが(例えば10より大きいというデータであれば、倍の20ぐらいと想定する等)、今回は不等号は無視し、standard_valueをそのまま阻害値として採用した。
- 複数の実験等によりCHEMBL IDが重複して含まれる場合、阻害値はこれらの平均をとった。
- 最終的な目的変数として、論文では阻害値のしきい値を10uMとしているため、10uMより小さい値は1 (Active)、大きい値は0 (Inactive)として出力した。
以下にこれらの処理をしたスクリプト断片を示す。
rows_by_id = defaultdict(list)
# csvを読み込み、CHEMBLID毎にrowをまとめる
with open(args.input, "r") as f:
reader = csv.DictReader(f)
for row in reader:
rows_by_id[row["compound_chembl_id"]].append(row)
# activeとみなす値
threshold = 10000
# CHEMBLID毎に値を処理しながら出力する
with open(args.output, "w") as f:
writer = csv.writer(f, lineterminator="\n")
writer.writerow(["chembl_id", "canonical_smiles", "value", "outcome"])
# CHEMBLID毎に値を処理
for id in rows_by_id:
# 合計を求める
total = 0.0
for row in rows_by_id[id]:
value = row["standard_value"]
total += float(value)
# 平均を求める
mean = total/len(rows_by_id[id])
print(f'{id},{mean}')
outcome = 0
if mean < threshold:
outcome = 1
writer.writerow([id, rows_by_id[id][0]["canonical_smiles"], mean, outcome])
このプログラムを実行した結果、Active/Inactiveの件数に1件の差はあったものの、論文と同じ91件の学習データが得られた。論文では取得手順は書かれていなかったので、このデータの収集がほぼ再現できたことが今回一番嬉しかった。
5.1.2 PDBからのデータ取得
次にPDBからの取得を行う。実は取得手順は以前に書いた記事 バイオ系公共データベースからスクレイピングにより機械学習データを収集するとほぼ同じである。唯一の違いは、メインプロテアーゼかどうかを判断するために、macromoleculeという項目も収集している点である。scrapyを使わない版の収集プログラムを以下に再掲する。
import requests
import json
import time
import lxml.html
import argparse
import csv
def get_ligand(ligand_id):
tmp_url = "https://www.rcsb.org" + ligand_id
response = requests.get(tmp_url)
if response.status_code != 200:
return response.status_code, []
html = response.text
root = lxml.html.fromstring(html)
#print(html)
print(tmp_url)
smiles = root.xpath("//tr[@id='chemicalIsomeric']/td[1]/text()")[0]
inchi = root.xpath("//tr[@id='chemicalInChI']/td[1]/text()")[0]
inchi_key = root.xpath("//tr[@id='chemicalInChIKey']/td[1]/text()")[0]
return response.status_code, [smiles, inchi, inchi_key]
def get_structure(structure_id):
structure_url = "https://www.rcsb.org/structure/"
tmp_url = structure_url + structure_id
print(tmp_url)
html = requests.get(tmp_url).text
root = lxml.html.fromstring(html)
macromolecule_trs = root.xpath("//tr[contains(@id,'macromolecule-entityId-')]")
macromolecule = ""
for tr in macromolecule_trs:
print(tr.xpath("@id"))
macromolecule += tr.xpath("td[position()=1]/text()")[0] + ","
print(f"macro={macromolecule}")
binding_trs = root.xpath("//tr[contains(@id,'binding_row')]")
datas = []
ids = []
for tr in binding_trs:
print(tr.xpath("@id"))
d1 = tr.xpath("td[position()=1]/a/@href")
if d1[0] in ids:
continue
ids.append(d1[0])
status_code, values = get_ligand(d1[0])
ligand_id = d1[0][(d1[0].rfind("/") + 1):]
print(ligand_id)
if status_code == 200:
smiles, inchi, inchi_key = values
#item = tr.xpath("a/td[position()=2]/text()")[0]
item = tr.xpath("td[position()=2]/a/text()")[0]
item = item.strip()
value = tr.xpath("td[position()=2]/text()")[0]
value = value.replace(":", "")
value = value.replace(";", "")
value = value.replace(" ", "")
value = value.replace("\n", "")
print(value)
values = value.split(" ", 1)
print(values)
value = values[0].strip()
unit = values[1].strip()
datas.append([ligand_id, smiles, inchi, inchi_key, item, value, unit, macromolecule])
time.sleep(1)
return datas
def main():
parser = argparse.ArgumentParser()
parser.add_argument("-output", type=str, required=True)
args = parser.parse_args()
base_url = "https://www.rcsb.org/search/data"
payloads = {"query":{"type":"group","logical_operator":"and","nodes":[{"type":"group","logical_operator":"and","nodes":[{"type":"group","logical_operator":"or","nodes":[{"type":"group","logical_operator":"and","nodes":[{"type":"terminal","service":"text","parameters":{"attribute":"rcsb_binding_affinity.value","negation":False,"operator":"exists"},"node_id":0},{"type":"terminal","service":"text","parameters":{"attribute":"rcsb_binding_affinity.type","operator":"exact_match","value":"IC50"},"node_id":1}],"label":"nested-attribute"},{"type":"group","logical_operator":"and","nodes":[{"type":"terminal","service":"text","parameters":{"attribute":"rcsb_binding_affinity.value","negation":False,"operator":"exists"},"node_id":2},{"type":"terminal","service":"text","parameters":{"attribute":"rcsb_binding_affinity.type","operator":"exact_match","value":"Ki"},"node_id":3}],"label":"nested-attribute"}]},{"type":"group","logical_operator":"and","nodes":[{"type":"terminal","service":"text","parameters":{"operator":"exact_match","negation":False,"value":"Severe acute respiratory syndrome-related coronavirus","attribute":"rcsb_entity_source_organism.taxonomy_lineage.name"},"node_id":4}]}],"label":"text"}],"label":"query-builder"},"return_type":"entry","request_options":{"pager":{"start":0,"rows":100},"scoring_strategy":"combined","sort":[{"sort_by":"score","direction":"desc"}]},"request_info":{"src":"ui","query_id":"e757fdfd5f9fb0efa272769c5966e3f4"}}
print(json.dumps(payloads))
response = requests.post(
base_url,
json.dumps(payloads),
headers={'Content-Type': 'application/json'})
datas = []
for a in response.json()["result_set"]:
structure_id = a["identifier"]
datas.extend(get_structure(structure_id))
time.sleep(1)
with open(args.output, "w") as f:
writer = csv.writer(f, lineterminator="\n")
writer.writerow(["ligand_id", "canonical_smiles", "inchi", "inchi_key", "item", "value", "unit"])
for data in datas:
writer.writerow(data)
if __name__ == "__main__":
main()
これを実行すると以下のような形式のデータが21件得られる。一番右の項目にタンパク質の名前があるので、"3C like proteinase"や、"main proteinase"といったフレーズが含まれていないものは除外し、最終的に10件を学習データとした。
ligand_id,canonical_smiles,inchi,inchi_key,item,value,unit
PMA,OC(=O)c1cc(C(O)=O)c(cc1C(O)=O)C(O)=O,"InChI=1S/C10H6O8/c11-7(12)3-1-4(8(13)14)6(10(17)18)2-5(3)9(15)16/h1-2H,(H,11,12)(H,13,14)(H,15,16)(H,17,18)",CYIDZMCFTVVTJO-UHFFFAOYSA-N,Ki,700,nM,"3C-like proteinase,"
TLD,Cc1ccc(S)c(S)c1,"InChI=1S/C7H8S2/c1-5-2-3-6(8)7(9)4-5/h2-4,8-9H,1H3",NIAAGQAEVGMHPM-UHFFFAOYSA-N,Ki,1400,nM,"Replicase polyprotein 1ab,"
ZU5,CC(C)C[C@H](NC(=O)[C@@H](NC(=O)OCc1ccccc1)[C@@H](C)OC(C)(C)C)C(=O)N[C@H](CCC(=O)C2CC2)C[C@@H]3CCNC3=O,"InChI=1S/C34H52N4O7/c1-21(2)18-27(31(41)36-26(14-15-28(39)24-12-13-24)19-25-16-17-35-30(25)40)37-32(42)29(22(3)45-34(4,5)6)38-33(43)44-20-23-10-8-7-9-11-23/h7-11,21-22,24-27,29H,12-20H2,1-6H3,(H,35,40)(H,36,41)(H,37,42)(H,38,43)/t22-,25+,26-,27+,29+/m1/s1",QIMPWBPEAHOISN-XSLDCGIXSA-N,Ki,99,nM,"3C-like proteinase,"
ZU3,CC(C)C[C@H](NC(=O)[C@H](CNC(=O)C(C)(C)C)NC(=O)OCc1ccccc1)C(=O)N[C@H](CCC(C)=O)C[C@@H]2CCNC2=O,"InChI=1S/C32H49N5O7/c1-20(2)16-25(28(40)35-24(13-12-21(3)38)17-23-14-15-33-27(23)39)36-29(41)26(18-34-30(42)32(4,5)6)37-31(43)44-19-22-10-8-7-9-11-22/h7-11,20,23-26H,12-19H2,1-6H3,(H,33,39)(H,34,42)(H,35,40)(H,36,41)(H,37,43)/t23-,24+,25-,26-/m0/s1",IEQRDAZPCPYZAJ-QYOOZWMWSA-N,Ki,38,nM,"3C-like proteinase,"
S89,OC[C@H](Cc1ccccc1)NC(=O)[C@H](Cc2ccccc2)NC(=O)/C=C/c3ccccc3,"InChI=1S/C27H28N2O3/c30-20-24(18-22-12-6-2-7-13-22)28-27(32)25(19-23-14-8-3-9-15-23)29-26(31)17-16-21-10-4-1-5-11-21/h1-17,24-25,30H,18-20H2,(H,28,32)(H,29,31)/b17-16+/t24-,25-/m0/s1",GEVQDXBVGFGWFA-KQRRRSJSSA-N,Ki,2240,nM,"3C-like proteinase,"
残念ながらChEMBLのようにはうまくいかず論文の22件は得られなかった。また上の10件の中には、論文の22件に含まれていないものも何件かあった。
これについては、次のいずれかまたは、組み合わせが原因と考えている。
- PDBに対する検索の指定の仕方に誤り、漏れがあった。
- タンパク質の構造に紐づけられているLingadデータのみを収集対象したが、実は紐づけられていないものについても、リンク先の論文の中にデータが存在していた可能性がある。
原因は不明だが、今回の指定手順で得られたデータ自体が間違っているとも思えなかったので、PDBについてはこれら10件を採用することとした。
5.1.3 ChEMBLとPDBのデータのマージ
5.1.1, 5.1.2で得られたデータをマージし、最終的に101件を学習データとしてcsvに出力した。スクリプトは省略する。
5.2 バーチャルスクリーニング用データの準備
本論文では、市販/中止/実験、および治験薬に対し、構築した予測モデルを適用している。このために必要となる、市販/中止/実験、および治験薬の構造データをDrugBankより取得しておく。論文のgithubには整形されたデータがそのままあるが、しつこいくらいに自分でやってみる。
5.2.1 DrugBankからのデータ取得
まずXMLデータとして、https://www.drugbank.ca/releases/latestの"DOWNLOAD (XML)"と記載のあるリンクから、xmlをダウンロードする。
続いてhttps://www.drugbank.ca/releases/latest#structuresからsdfをダウンロードする。
アカウントは事前に作成しておく必要があったかもしれない。
これらを"full database.xml", "open structures.sdf"として保存する。
5.2.2 DrugBankデータの整形
sdfに構造データが格納されておりこれだけでも予測モデルを適用することはできるが、xmlには市販/中止/実験等の種別があるため、これらを関連づけてcsvに保存する。
以下にスクリプト断片を示す。
def get_group(groups_node, ns):
ret = ""
if groups_node:
for i, child in enumerate(groups_node.iter(f"{ns}group")):
if i > 0 :
ret += ","
ret += child.text
return ret
def get_id(drug_node, ns):
for i, child in enumerate(drug_node.iter(f"{ns}drugbank-id")):
for attr in child.attrib:
if attr == "primary":
return child.text
return None
def main():
parser = argparse.ArgumentParser()
parser.add_argument("-input_xml", type=str, required=True)
parser.add_argument("-input_sdf", type=str, required=True)
parser.add_argument("-output", type=str, required=True)
args = parser.parse_args()
name_dict = {}
smiles_dict = {}
sdf_sup = Chem.SDMolSupplier(args.input_sdf)
datas = []
for i, mol in enumerate(sdf_sup):
if not mol:
continue
if mol.HasProp("DRUGBANK_ID"):
id = mol.GetProp("DRUGBANK_ID")
if mol.HasProp("COMMON_NAME"):
name = mol.GetProp("COMMON_NAME")
smiles = Chem.MolToSmiles(mol)
name_dict[id] = name
smiles_dict[id] = smiles
print(f"{i} {id} {name} {smiles}")
tree = ET.parse(args.input_xml)
root = tree.getroot()
ns = "{http://www.drugbank.ca}"
ids = []
datas = []
for i, drug in enumerate(root.iter(f"{ns}drug")):
id = get_id(drug, ns)
category = get_group(drug.find(f"{ns}groups"), ns)
if id and id in smiles_dict:
print(f"{i}, {id}, {category}")
ids.append(id)
datas.append([name_dict[id], category, smiles_dict[id]])
df = pd.DataFrame(datas, index=ids, columns=[["name", "status", "smiles"]])
df.to_csv(args.output)
if __name__ == "__main__":
main()
このスクリプトを実行して得られるデータはこんな感じになる。構造データはSMILESとして保存している。
id,name,status,smiles
DB00006,Bivalirudin,"approved,investigational",CC[C@H](C)[C@H](NC(=O)[C@H](CCC(=O)O)NC(=O)[C@H](CCC(=O)O)NC(=O)[C@H](Cc1ccccc1)NC(=O)[C@H](CC(=O)O)NC(=O)CNC(=O)[C@H](CC(N)=O)NC(=O)CNC(=O)CNC(=O)CNC(=O)CNC(=O)[C@@H]1CCCN1C(=O)[C@H](CCCNC(=N)N)NC(=O)[C@@H]1CCCN1C(=O)[C@H](N)Cc1ccccc1)C(=O)N1CCC[C@H]1C(=O)N[C@@H](CCC(=O)O)C(=O)N[C@@H](CCC(=O)O)C(=O)N[C@@H](Cc1ccc(O)cc1)C(=O)N[C@@H](CC(C)C)C(=O)O
DB00007,Leuprolide,"approved,investigational",CCNC(=O)[C@@H]1CCCN1C(=O)[C@H](CCCNC(=N)N)NC(=O)[C@H](CC(C)C)NC(=O)[C@@H](CC(C)C)NC(=O)[C@H](Cc1ccc(O)cc1)NC(=O)[C@H](CO)NC(=O)[C@H](Cc1c[nH]c2ccccc12)NC(=O)[C@H](Cc1c[nH]cn1)NC(=O)[C@@H]1CCC(=O)N1
DB00014,Goserelin,approved,CC(C)C[C@H](NC(=O)[C@@H](COC(C)(C)C)NC(=O)[C@H](Cc1ccc(O)cc1)NC(=O)[C@H](CO)NC(=O)[C@H](Cc1c[nH]c2ccccc12)NC(=O)[C@H](Cc1cnc[nH]1)NC(=O)[C@@H]1CCC(=O)N1)C(=O)N[C@@H](CCCN=C(N)N)C(=O)N1CCC[C@H]1C(=O)NNC(N)=O
DB00027,Gramicidin D,approved,CC(C)C[C@@H](NC(=O)CNC(=O)[C@@H](NC=O)C(C)C)C(=O)N[C@@H](C)C(=O)N[C@@H](C(=O)N[C@H](C(=O)N[C@@H](C(=O)N[C@@H](Cc1c[nH]c2ccccc12)C(=O)N[C@H](CC(C)C)C(=O)N[C@@H](Cc1c[nH]c2ccccc12)C(=O)N[C@H](CC(C)C)C(=O)N[C@@H](Cc1c[nH]c2ccccc12)C(=O)N[C@H](CC(C)C)C(=O)N[C@@H](Cc1c[nH]c2ccccc12)C(=O)NCCO)C(C)C)C(C)C)C(C)C
DB00035,Desmopressin,approved,N=C(N)NCCC[C@@H](NC(=O)[C@@H]1CCCN1C(=O)[C@@H]1CSSCCC(=O)N[C@@H](Cc2ccc(O)cc2)C(=O)N[C@@H](Cc2ccccc2)C(=O)N[C@@H](CCC(N)=O)C(=O)N[C@@H](CC(N)=O)C(=O)N1)C(=O)NCC(N)=O
5.3 予測モデルの構築
次に学習データを使って予測モデルを構築する。
5.3.1 構造データからのフィンガープリント生成
ここでは、5.1.3 で生成されたcsvファイルから機械学習用の説明変数を生成する。
具体的には、csvファイルの"canonical_smiles"列にあるSMILESを読み取り、RDKitを使ってmolオブジェクトに変換する。その後、RDKitのAllChem.GetMorganFingerprintAsBitVectを使って2048ビットのフィンガープリントを生成する。フィンガープリントとはざっくり説明すると、化合物に特定の構造が含まれていれば、それに対応したビットが1、そうでない場合は0となるものがいくつかか集まったものである。今回は2048個の0または1で構成されるビット列が生成される。
from rdkit import Chem
from rdkit.Chem import AllChem
from molvs.normalize import Normalizer, Normalization
from molvs.tautomer import TAUTOMER_TRANSFORMS, TAUTOMER_SCORES, MAX_TAUTOMERS, TautomerCanonicalizer, TautomerEnumerator, TautomerTransform
from molvs.fragment import LargestFragmentChooser
from molvs.charge import Reionizer, Uncharger
import argparse
import csv
import pandas as pd
import numpy as np
def normalize(smiles):
# Generate Mol
mol = Chem.MolFromSmiles(smiles)
# Uncharge
uncharger = Uncharger()
mol = uncharger.uncharge(mol)
# LargestFragmentChooser
flagmentChooser = LargestFragmentChooser()
mol = flagmentChooser(mol)
# Sanitaize
Chem.SanitizeMol(mol)
# Normalize
normalizer = Normalizer()
mol = normalizer.normalize(mol)
tautomerCanonicalizer = TautomerCanonicalizer()
mol = tautomerCanonicalizer.canonicalize(mol)
return Chem.MolToSmiles(mol)
def main():
parser = argparse.ArgumentParser()
parser.add_argument("-input", type=str, required=True)
parser.add_argument("-output", type=str, required=True)
args = parser.parse_args()
#学習データの読み込み
smiles_list = list()
datas = []
with open(args.input, "r") as f:
reader = csv.DictReader(f)
for row in reader:
org_smiles = row["canonical_smiles"]
new_smiles = normalize(org_smiles)
existFlag = False
for tmp_smiles in smiles_list:
if new_smiles == tmp_smiles:
print("exist!")
existFlag = True
break
if not existFlag:
smiles_list.append(new_smiles)
mol = Chem.MolFromSmiles(new_smiles)
fp = AllChem.GetMorganFingerprintAsBitVect(mol, radius=3, nBits=2048, useFeatures=False, useChirality=False)
fp = pd.Series(np.asarray(fp)).values
data = []
data.append(row["id"])
data.append(int(row["outcome"]))
data.extend(fp)
datas.append(data)
columns = list()
columns.append("id")
columns.append("outcome")
columns.extend(["Bit_" + str(i + 1) for i in range(2048)])
df = pd.DataFrame(data=datas, columns=columns)
df.set_index("id", inplace=True, drop=True)
#保存
df.to_csv(args.output)
if __name__ == "__main__":
main()
化合物の前処理、標準化を行い、念のため同じ化合物が存在した場合に除去するという処理も入れた。化合物から3次元座標を生成するという処理も行われことがあるが、今回利用したフィンガープリントの場合3次元座標は不要のため、処理はしていない。
実行後すると以下のようなデータが得られる。1列目がID、2列目が目的変数、3列目以降がフィンガープリントによる説明変数である。
id,outcome,Bit_1,Bit_2,Bit_3,Bit_4,Bit_5,Bit_6,Bit_7,Bit_8,Bit_9,Bit_10,Bit_11,Bit_12,Bit_13,Bit_14,Bit_15,Bit_16,Bit_17,Bit_18,Bit_19,Bit_20,Bit_21,Bit_22,Bit_23,Bit_24,Bit_25,Bit_26,Bit_27,Bit_28,Bit_29,Bit_30,Bit_31,Bit_32,Bit_33,Bit_34,Bit_35,Bit_36,Bit_37,Bit_38,Bit_39,Bit_40,Bit_41,Bit_42,Bit_43,Bit_44,Bit_45,Bit_46,Bit_47,Bit_48,Bit_49,Bit_50,Bit_51,Bit_52,Bit_53,Bit_54,Bit_55,Bit_56,Bit_57,Bit_58,Bit_59,Bit_60,Bit_61,Bit_62,Bit_63,Bit_64,Bit_65,Bit_66,Bit_67,Bit_68,Bit_69,Bit_70,Bit_71,Bit_72,Bit_73,Bit_74,Bit_75,Bit_76,Bit_77,Bit_78,Bit_79,Bit_80,Bit_81,Bit_82,Bit_83,Bit_84,Bit_85,Bit_86,Bit_87,Bit_88,Bit_89,Bit_90,Bit_91,Bit_92,Bit_93,Bit_94,Bit_95,Bit_96,Bit_97,Bit_98,Bit_99,Bit_100,Bit_101,Bit_102,Bit_103,Bit_104,Bit_105,Bit_106,Bit_107,Bit_108,Bit_109,Bit_110,Bit_111,Bit_112,Bit_113,Bit_114,Bit_115,Bit_116,Bit_117,Bit_118,Bit_119,Bit_120,Bit_121,Bit_122,Bit_123,Bit_124,Bit_125,Bit_126,Bit_127,Bit_128,Bit_129,Bit_130,Bit_131,Bit_132,Bit_133,Bit_134,Bit_135,Bit_136,Bit_137,Bit_138,Bit_139,Bit_140,Bit_141,Bit_142,Bit_143,Bit_144,Bit_145,Bit_146,Bit_147,Bit_148,Bit_149,Bit_150,Bit_151,Bit_152,Bit_153,Bit_154,Bit_155,Bit_156,Bit_157,Bit_158,Bit_159,Bit_160,Bit_161,Bit_162,Bit_163,Bit_164,Bit_165,Bit_166,Bit_167,Bit_168,Bit_169,Bit_170,Bit_171,Bit_172,Bit_173,Bit_174,Bit_175,Bit_176,Bit_177,Bit_178,Bit_179,Bit_180,Bit_181,Bit_182,Bit_183,Bit_184,Bit_185,Bit_186,Bit_187,Bit_188,Bit_189,Bit_190,Bit_191,Bit_192,Bit_193,Bit_194,Bit_195,Bit_196,Bit_197,Bit_198,Bit_199,Bit_200,Bit_201,Bit_202,Bit_203,Bit_204,Bit_205,Bit_206,Bit_207,Bit_208,Bit_209,Bit_210,Bit_211,Bit_212,Bit_213,Bit_214,Bit_215,Bit_216,Bit_217,Bit_218,Bit_219,Bit_220,Bit_221,Bit_222,Bit_223,Bit_224,Bit_225,Bit_226,Bit_227,Bit_228,Bit_229,Bit_230,Bit_231,Bit_232,Bit_233,Bit_234,Bit_235,Bit_236,Bit_237,Bit_238,Bit_239,Bit_240,Bit_241,Bit_242,Bit_243,Bit_244,Bit_245,Bit_246,Bit_247,Bit_248,Bit_249,Bit_250,Bit_251,Bit_252,Bit_253,Bit_254,Bit_255,Bit_256,Bit_257,Bit_258,Bit_259,Bit_260,Bit_261,Bit_262,Bit_263,Bit_264,Bit_265,Bit_266,Bit_267,Bit_268,Bit_269,Bit_270,Bit_271,Bit_272,Bit_273,Bit_274,Bit_275,Bit_276,Bit_277,Bit_278,Bit_279,Bit_280,Bit_281,Bit_282,Bit_283,Bit_284,Bit_285,Bit_286,Bit_287,Bit_288,Bit_289,Bit_290,Bit_291,Bit_292,Bit_293,Bit_294,Bit_295,Bit_296,Bit_297,Bit_298,Bit_299,Bit_300,Bit_301,Bit_302,Bit_303,Bit_304,Bit_305,Bit_306,Bit_307,Bit_308,Bit_309,Bit_310,Bit_311,Bit_312,Bit_313,Bit_314,Bit_315,Bit_316,Bit_317,Bit_318,Bit_319,Bit_320,Bit_321,Bit_322,Bit_323,Bit_324,Bit_325,Bit_326,Bit_327,Bit_328,Bit_329,Bit_330,Bit_331,Bit_332,Bit_333,Bit_334,Bit_335,Bit_336,Bit_337,Bit_338,Bit_339,Bit_340,Bit_341,Bit_342,Bit_343,Bit_344,Bit_345,Bit_346,Bit_347,Bit_348,Bit_349,Bit_350,Bit_351,Bit_352,Bit_353,Bit_354,Bit_355,Bit_356,Bit_357,Bit_358,Bit_359,Bit_360,Bit_361,Bit_362,Bit_363,Bit_364,Bit_365,Bit_366,Bit_367,Bit_368,Bit_369,Bit_370,Bit_371,Bit_372,Bit_373,Bit_374,Bit_375,Bit_376,Bit_377,Bit_378,Bit_379,Bit_380,Bit_381,Bit_382,Bit_383,Bit_384,Bit_385,Bit_386,Bit_387,Bit_388,Bit_389,Bit_390,Bit_391,Bit_392,Bit_393,Bit_394,Bit_395,Bit_396,Bit_397,Bit_398,Bit_399,Bit_400,Bit_401,Bit_402,Bit_403,Bit_404,Bit_405,Bit_406,Bit_407,Bit_408,Bit_409,Bit_410,Bit_411,Bit_412,Bit_413,Bit_414,Bit_415,Bit_416,Bit_417,Bit_418,Bit_419,Bit_420,Bit_421,Bit_422,Bit_423,Bit_424,Bit_425,Bit_426,Bit_427,Bit_428,Bit_429,Bit_430,Bit_431,Bit_432,Bit_433,Bit_434,Bit_435,Bit_436,Bit_437,Bit_438,Bit_439,Bit_440,Bit_441,Bit_442,Bit_443,Bit_444,Bit_445,Bit_446,Bit_447,Bit_448,Bit_449,Bit_450,Bit_451,Bit_452,Bit_453,Bit_454,Bit_455,Bit_456,Bit_457,Bit_458,Bit_459,Bit_460,Bit_461,Bit_462,Bit_463,Bit_464,Bit_465,Bit_466,Bit_467,Bit_468,Bit_469,Bit_470,Bit_471,Bit_472,Bit_473,Bit_474,Bit_475,Bit_476,Bit_477,Bit_478,Bit_479,Bit_480,Bit_481,Bit_482,Bit_483,Bit_484,Bit_485,Bit_486,Bit_487,Bit_488,Bit_489,Bit_490,Bit_491,Bit_492,Bit_493,Bit_494,Bit_495,Bit_496,Bit_497,Bit_498,Bit_499,Bit_500,Bit_501,Bit_502,Bit_503,Bit_504,Bit_505,Bit_506,Bit_507,Bit_508,Bit_509,Bit_510,Bit_511,Bit_512,Bit_513,Bit_514,Bit_515,Bit_516,Bit_517,Bit_518,Bit_519,Bit_520,Bit_521,Bit_522,Bit_523,Bit_524,Bit_525,Bit_526,Bit_527,Bit_528,Bit_529,Bit_530,Bit_531,Bit_532,Bit_533,Bit_534,Bit_535,Bit_536,Bit_537,Bit_538,Bit_539,Bit_540,Bit_541,Bit_542,Bit_543,Bit_544,Bit_545,Bit_546,Bit_547,Bit_548,Bit_549,Bit_550,Bit_551,Bit_552,Bit_553,Bit_554,Bit_555,Bit_556,Bit_557,Bit_558,Bit_559,Bit_560,Bit_561,Bit_562,Bit_563,Bit_564,Bit_565,Bit_566,Bit_567,Bit_568,Bit_569,Bit_570,Bit_571,Bit_572,Bit_573,Bit_574,Bit_575,Bit_576,Bit_577,Bit_578,Bit_579,Bit_580,Bit_581,Bit_582,Bit_583,Bit_584,Bit_585,Bit_586,Bit_587,Bit_588,Bit_589,Bit_590,Bit_591,Bit_592,Bit_593,Bit_594,Bit_595,Bit_596,Bit_597,Bit_598,Bit_599,Bit_600,Bit_601,Bit_602,Bit_603,Bit_604,Bit_605,Bit_606,Bit_607,Bit_608,Bit_609,Bit_610,Bit_611,Bit_612,Bit_613,Bit_614,Bit_615,Bit_616,Bit_617,Bit_618,Bit_619,Bit_620,Bit_621,Bit_622,Bit_623,Bit_624,Bit_625,Bit_626,Bit_627,Bit_628,Bit_629,Bit_630,Bit_631,Bit_632,Bit_633,Bit_634,Bit_635,Bit_636,Bit_637,Bit_638,Bit_639,Bit_640,Bit_641,Bit_642,Bit_643,Bit_644,Bit_645,Bit_646,Bit_647,Bit_648,Bit_649,Bit_650,Bit_651,Bit_652,Bit_653,Bit_654,Bit_655,Bit_656,Bit_657,Bit_658,Bit_659,Bit_660,Bit_661,Bit_662,Bit_663,Bit_664,Bit_665,Bit_666,Bit_667,Bit_668,Bit_669,Bit_670,Bit_671,Bit_672,Bit_673,Bit_674,Bit_675,Bit_676,Bit_677,Bit_678,Bit_679,Bit_680,Bit_681,Bit_682,Bit_683,Bit_684,Bit_685,Bit_686,Bit_687,Bit_688,Bit_689,Bit_690,Bit_691,Bit_692,Bit_693,Bit_694,Bit_695,Bit_696,Bit_697,Bit_698,Bit_699,Bit_700,Bit_701,Bit_702,Bit_703,Bit_704,Bit_705,Bit_706,Bit_707,Bit_708,Bit_709,Bit_710,Bit_711,Bit_712,Bit_713,Bit_714,Bit_715,Bit_716,Bit_717,Bit_718,Bit_719,Bit_720,Bit_721,Bit_722,Bit_723,Bit_724,Bit_725,Bit_726,Bit_727,Bit_728,Bit_729,Bit_730,Bit_731,Bit_732,Bit_733,Bit_734,Bit_735,Bit_736,Bit_737,Bit_738,Bit_739,Bit_740,Bit_741,Bit_742,Bit_743,Bit_744,Bit_745,Bit_746,Bit_747,Bit_748,Bit_749,Bit_750,Bit_751,Bit_752,Bit_753,Bit_754,Bit_755,Bit_756,Bit_757,Bit_758,Bit_759,Bit_760,Bit_761,Bit_762,Bit_763,Bit_764,Bit_765,Bit_766,Bit_767,Bit_768,Bit_769,Bit_770,Bit_771,Bit_772,Bit_773,Bit_774,Bit_775,Bit_776,Bit_777,Bit_778,Bit_779,Bit_780,Bit_781,Bit_782,Bit_783,Bit_784,Bit_785,Bit_786,Bit_787,Bit_788,Bit_789,Bit_790,Bit_791,Bit_792,Bit_793,Bit_794,Bit_795,Bit_796,Bit_797,Bit_798,Bit_799,Bit_800,Bit_801,Bit_802,Bit_803,Bit_804,Bit_805,Bit_806,Bit_807,Bit_808,Bit_809,Bit_810,Bit_811,Bit_812,Bit_813,Bit_814,Bit_815,Bit_816,Bit_817,Bit_818,Bit_819,Bit_820,Bit_821,Bit_822,Bit_823,Bit_824,Bit_825,Bit_826,Bit_827,Bit_828,Bit_829,Bit_830,Bit_831,Bit_832,Bit_833,Bit_834,Bit_835,Bit_836,Bit_837,Bit_838,Bit_839,Bit_840,Bit_841,Bit_842,Bit_843,Bit_844,Bit_845,Bit_846,Bit_847,Bit_848,Bit_849,Bit_850,Bit_851,Bit_852,Bit_853,Bit_854,Bit_855,Bit_856,Bit_857,Bit_858,Bit_859,Bit_860,Bit_861,Bit_862,Bit_863,Bit_864,Bit_865,Bit_866,Bit_867,Bit_868,Bit_869,Bit_870,Bit_871,Bit_872,Bit_873,Bit_874,Bit_875,Bit_876,Bit_877,Bit_878,Bit_879,Bit_880,Bit_881,Bit_882,Bit_883,Bit_884,Bit_885,Bit_886,Bit_887,Bit_888,Bit_889,Bit_890,Bit_891,Bit_892,Bit_893,Bit_894,Bit_895,Bit_896,Bit_897,Bit_898,Bit_899,Bit_900,Bit_901,Bit_902,Bit_903,Bit_904,Bit_905,Bit_906,Bit_907,Bit_908,Bit_909,Bit_910,Bit_911,Bit_912,Bit_913,Bit_914,Bit_915,Bit_916,Bit_917,Bit_918,Bit_919,Bit_920,Bit_921,Bit_922,Bit_923,Bit_924,Bit_925,Bit_926,Bit_927,Bit_928,Bit_929,Bit_930,Bit_931,Bit_932,Bit_933,Bit_934,Bit_935,Bit_936,Bit_937,Bit_938,Bit_939,Bit_940,Bit_941,Bit_942,Bit_943,Bit_944,Bit_945,Bit_946,Bit_947,Bit_948,Bit_949,Bit_950,Bit_951,Bit_952,Bit_953,Bit_954,Bit_955,Bit_956,Bit_957,Bit_958,Bit_959,Bit_960,Bit_961,Bit_962,Bit_963,Bit_964,Bit_965,Bit_966,Bit_967,Bit_968,Bit_969,Bit_970,Bit_971,Bit_972,Bit_973,Bit_974,Bit_975,Bit_976,Bit_977,Bit_978,Bit_979,Bit_980,Bit_981,Bit_982,Bit_983,Bit_984,Bit_985,Bit_986,Bit_987,Bit_988,Bit_989,Bit_990,Bit_991,Bit_992,Bit_993,Bit_994,Bit_995,Bit_996,Bit_997,Bit_998,Bit_999,Bit_1000,Bit_1001,Bit_1002,Bit_1003,Bit_1004,Bit_1005,Bit_1006,Bit_1007,Bit_1008,Bit_1009,Bit_1010,Bit_1011,Bit_1012,Bit_1013,Bit_1014,Bit_1015,Bit_1016,Bit_1017,Bit_1018,Bit_1019,Bit_1020,Bit_1021,Bit_1022,Bit_1023,Bit_1024,Bit_1025,Bit_1026,Bit_1027,Bit_1028,Bit_1029,Bit_1030,Bit_1031,Bit_1032,Bit_1033,Bit_1034,Bit_1035,Bit_1036,Bit_1037,Bit_1038,Bit_1039,Bit_1040,Bit_1041,Bit_1042,Bit_1043,Bit_1044,Bit_1045,Bit_1046,Bit_1047,Bit_1048,Bit_1049,Bit_1050,Bit_1051,Bit_1052,Bit_1053,Bit_1054,Bit_1055,Bit_1056,Bit_1057,Bit_1058,Bit_1059,Bit_1060,Bit_1061,Bit_1062,Bit_1063,Bit_1064,Bit_1065,Bit_1066,Bit_1067,Bit_1068,Bit_1069,Bit_1070,Bit_1071,Bit_1072,Bit_1073,Bit_1074,Bit_1075,Bit_1076,Bit_1077,Bit_1078,Bit_1079,Bit_1080,Bit_1081,Bit_1082,Bit_1083,Bit_1084,Bit_1085,Bit_1086,Bit_1087,Bit_1088,Bit_1089,Bit_1090,Bit_1091,Bit_1092,Bit_1093,Bit_1094,Bit_1095,Bit_1096,Bit_1097,Bit_1098,Bit_1099,Bit_1100,Bit_1101,Bit_1102,Bit_1103,Bit_1104,Bit_1105,Bit_1106,Bit_1107,Bit_1108,Bit_1109,Bit_1110,Bit_1111,Bit_1112,Bit_1113,Bit_1114,Bit_1115,Bit_1116,Bit_1117,Bit_1118,Bit_1119,Bit_1120,Bit_1121,Bit_1122,Bit_1123,Bit_1124,Bit_1125,Bit_1126,Bit_1127,Bit_1128,Bit_1129,Bit_1130,Bit_1131,Bit_1132,Bit_1133,Bit_1134,Bit_1135,Bit_1136,Bit_1137,Bit_1138,Bit_1139,Bit_1140,Bit_1141,Bit_1142,Bit_1143,Bit_1144,Bit_1145,Bit_1146,Bit_1147,Bit_1148,Bit_1149,Bit_1150,Bit_1151,Bit_1152,Bit_1153,Bit_1154,Bit_1155,Bit_1156,Bit_1157,Bit_1158,Bit_1159,Bit_1160,Bit_1161,Bit_1162,Bit_1163,Bit_1164,Bit_1165,Bit_1166,Bit_1167,Bit_1168,Bit_1169,Bit_1170,Bit_1171,Bit_1172,Bit_1173,Bit_1174,Bit_1175,Bit_1176,Bit_1177,Bit_1178,Bit_1179,Bit_1180,Bit_1181,Bit_1182,Bit_1183,Bit_1184,Bit_1185,Bit_1186,Bit_1187,Bit_1188,Bit_1189,Bit_1190,Bit_1191,Bit_1192,Bit_1193,Bit_1194,Bit_1195,Bit_1196,Bit_1197,Bit_1198,Bit_1199,Bit_1200,Bit_1201,Bit_1202,Bit_1203,Bit_1204,Bit_1205,Bit_1206,Bit_1207,Bit_1208,Bit_1209,Bit_1210,Bit_1211,Bit_1212,Bit_1213,Bit_1214,Bit_1215,Bit_1216,Bit_1217,Bit_1218,Bit_1219,Bit_1220,Bit_1221,Bit_1222,Bit_1223,Bit_1224,Bit_1225,Bit_1226,Bit_1227,Bit_1228,Bit_1229,Bit_1230,Bit_1231,Bit_1232,Bit_1233,Bit_1234,Bit_1235,Bit_1236,Bit_1237,Bit_1238,Bit_1239,Bit_1240,Bit_1241,Bit_1242,Bit_1243,Bit_1244,Bit_1245,Bit_1246,Bit_1247,Bit_1248,Bit_1249,Bit_1250,Bit_1251,Bit_1252,Bit_1253,Bit_1254,Bit_1255,Bit_1256,Bit_1257,Bit_1258,Bit_1259,Bit_1260,Bit_1261,Bit_1262,Bit_1263,Bit_1264,Bit_1265,Bit_1266,Bit_1267,Bit_1268,Bit_1269,Bit_1270,Bit_1271,Bit_1272,Bit_1273,Bit_1274,Bit_1275,Bit_1276,Bit_1277,Bit_1278,Bit_1279,Bit_1280,Bit_1281,Bit_1282,Bit_1283,Bit_1284,Bit_1285,Bit_1286,Bit_1287,Bit_1288,Bit_1289,Bit_1290,Bit_1291,Bit_1292,Bit_1293,Bit_1294,Bit_1295,Bit_1296,Bit_1297,Bit_1298,Bit_1299,Bit_1300,Bit_1301,Bit_1302,Bit_1303,Bit_1304,Bit_1305,Bit_1306,Bit_1307,Bit_1308,Bit_1309,Bit_1310,Bit_1311,Bit_1312,Bit_1313,Bit_1314,Bit_1315,Bit_1316,Bit_1317,Bit_1318,Bit_1319,Bit_1320,Bit_1321,Bit_1322,Bit_1323,Bit_1324,Bit_1325,Bit_1326,Bit_1327,Bit_1328,Bit_1329,Bit_1330,Bit_1331,Bit_1332,Bit_1333,Bit_1334,Bit_1335,Bit_1336,Bit_1337,Bit_1338,Bit_1339,Bit_1340,Bit_1341,Bit_1342,Bit_1343,Bit_1344,Bit_1345,Bit_1346,Bit_1347,Bit_1348,Bit_1349,Bit_1350,Bit_1351,Bit_1352,Bit_1353,Bit_1354,Bit_1355,Bit_1356,Bit_1357,Bit_1358,Bit_1359,Bit_1360,Bit_1361,Bit_1362,Bit_1363,Bit_1364,Bit_1365,Bit_1366,Bit_1367,Bit_1368,Bit_1369,Bit_1370,Bit_1371,Bit_1372,Bit_1373,Bit_1374,Bit_1375,Bit_1376,Bit_1377,Bit_1378,Bit_1379,Bit_1380,Bit_1381,Bit_1382,Bit_1383,Bit_1384,Bit_1385,Bit_1386,Bit_1387,Bit_1388,Bit_1389,Bit_1390,Bit_1391,Bit_1392,Bit_1393,Bit_1394,Bit_1395,Bit_1396,Bit_1397,Bit_1398,Bit_1399,Bit_1400,Bit_1401,Bit_1402,Bit_1403,Bit_1404,Bit_1405,Bit_1406,Bit_1407,Bit_1408,Bit_1409,Bit_1410,Bit_1411,Bit_1412,Bit_1413,Bit_1414,Bit_1415,Bit_1416,Bit_1417,Bit_1418,Bit_1419,Bit_1420,Bit_1421,Bit_1422,Bit_1423,Bit_1424,Bit_1425,Bit_1426,Bit_1427,Bit_1428,Bit_1429,Bit_1430,Bit_1431,Bit_1432,Bit_1433,Bit_1434,Bit_1435,Bit_1436,Bit_1437,Bit_1438,Bit_1439,Bit_1440,Bit_1441,Bit_1442,Bit_1443,Bit_1444,Bit_1445,Bit_1446,Bit_1447,Bit_1448,Bit_1449,Bit_1450,Bit_1451,Bit_1452,Bit_1453,Bit_1454,Bit_1455,Bit_1456,Bit_1457,Bit_1458,Bit_1459,Bit_1460,Bit_1461,Bit_1462,Bit_1463,Bit_1464,Bit_1465,Bit_1466,Bit_1467,Bit_1468,Bit_1469,Bit_1470,Bit_1471,Bit_1472,Bit_1473,Bit_1474,Bit_1475,Bit_1476,Bit_1477,Bit_1478,Bit_1479,Bit_1480,Bit_1481,Bit_1482,Bit_1483,Bit_1484,Bit_1485,Bit_1486,Bit_1487,Bit_1488,Bit_1489,Bit_1490,Bit_1491,Bit_1492,Bit_1493,Bit_1494,Bit_1495,Bit_1496,Bit_1497,Bit_1498,Bit_1499,Bit_1500,Bit_1501,Bit_1502,Bit_1503,Bit_1504,Bit_1505,Bit_1506,Bit_1507,Bit_1508,Bit_1509,Bit_1510,Bit_1511,Bit_1512,Bit_1513,Bit_1514,Bit_1515,Bit_1516,Bit_1517,Bit_1518,Bit_1519,Bit_1520,Bit_1521,Bit_1522,Bit_1523,Bit_1524,Bit_1525,Bit_1526,Bit_1527,Bit_1528,Bit_1529,Bit_1530,Bit_1531,Bit_1532,Bit_1533,Bit_1534,Bit_1535,Bit_1536,Bit_1537,Bit_1538,Bit_1539,Bit_1540,Bit_1541,Bit_1542,Bit_1543,Bit_1544,Bit_1545,Bit_1546,Bit_1547,Bit_1548,Bit_1549,Bit_1550,Bit_1551,Bit_1552,Bit_1553,Bit_1554,Bit_1555,Bit_1556,Bit_1557,Bit_1558,Bit_1559,Bit_1560,Bit_1561,Bit_1562,Bit_1563,Bit_1564,Bit_1565,Bit_1566,Bit_1567,Bit_1568,Bit_1569,Bit_1570,Bit_1571,Bit_1572,Bit_1573,Bit_1574,Bit_1575,Bit_1576,Bit_1577,Bit_1578,Bit_1579,Bit_1580,Bit_1581,Bit_1582,Bit_1583,Bit_1584,Bit_1585,Bit_1586,Bit_1587,Bit_1588,Bit_1589,Bit_1590,Bit_1591,Bit_1592,Bit_1593,Bit_1594,Bit_1595,Bit_1596,Bit_1597,Bit_1598,Bit_1599,Bit_1600,Bit_1601,Bit_1602,Bit_1603,Bit_1604,Bit_1605,Bit_1606,Bit_1607,Bit_1608,Bit_1609,Bit_1610,Bit_1611,Bit_1612,Bit_1613,Bit_1614,Bit_1615,Bit_1616,Bit_1617,Bit_1618,Bit_1619,Bit_1620,Bit_1621,Bit_1622,Bit_1623,Bit_1624,Bit_1625,Bit_1626,Bit_1627,Bit_1628,Bit_1629,Bit_1630,Bit_1631,Bit_1632,Bit_1633,Bit_1634,Bit_1635,Bit_1636,Bit_1637,Bit_1638,Bit_1639,Bit_1640,Bit_1641,Bit_1642,Bit_1643,Bit_1644,Bit_1645,Bit_1646,Bit_1647,Bit_1648,Bit_1649,Bit_1650,Bit_1651,Bit_1652,Bit_1653,Bit_1654,Bit_1655,Bit_1656,Bit_1657,Bit_1658,Bit_1659,Bit_1660,Bit_1661,Bit_1662,Bit_1663,Bit_1664,Bit_1665,Bit_1666,Bit_1667,Bit_1668,Bit_1669,Bit_1670,Bit_1671,Bit_1672,Bit_1673,Bit_1674,Bit_1675,Bit_1676,Bit_1677,Bit_1678,Bit_1679,Bit_1680,Bit_1681,Bit_1682,Bit_1683,Bit_1684,Bit_1685,Bit_1686,Bit_1687,Bit_1688,Bit_1689,Bit_1690,Bit_1691,Bit_1692,Bit_1693,Bit_1694,Bit_1695,Bit_1696,Bit_1697,Bit_1698,Bit_1699,Bit_1700,Bit_1701,Bit_1702,Bit_1703,Bit_1704,Bit_1705,Bit_1706,Bit_1707,Bit_1708,Bit_1709,Bit_1710,Bit_1711,Bit_1712,Bit_1713,Bit_1714,Bit_1715,Bit_1716,Bit_1717,Bit_1718,Bit_1719,Bit_1720,Bit_1721,Bit_1722,Bit_1723,Bit_1724,Bit_1725,Bit_1726,Bit_1727,Bit_1728,Bit_1729,Bit_1730,Bit_1731,Bit_1732,Bit_1733,Bit_1734,Bit_1735,Bit_1736,Bit_1737,Bit_1738,Bit_1739,Bit_1740,Bit_1741,Bit_1742,Bit_1743,Bit_1744,Bit_1745,Bit_1746,Bit_1747,Bit_1748,Bit_1749,Bit_1750,Bit_1751,Bit_1752,Bit_1753,Bit_1754,Bit_1755,Bit_1756,Bit_1757,Bit_1758,Bit_1759,Bit_1760,Bit_1761,Bit_1762,Bit_1763,Bit_1764,Bit_1765,Bit_1766,Bit_1767,Bit_1768,Bit_1769,Bit_1770,Bit_1771,Bit_1772,Bit_1773,Bit_1774,Bit_1775,Bit_1776,Bit_1777,Bit_1778,Bit_1779,Bit_1780,Bit_1781,Bit_1782,Bit_1783,Bit_1784,Bit_1785,Bit_1786,Bit_1787,Bit_1788,Bit_1789,Bit_1790,Bit_1791,Bit_1792,Bit_1793,Bit_1794,Bit_1795,Bit_1796,Bit_1797,Bit_1798,Bit_1799,Bit_1800,Bit_1801,Bit_1802,Bit_1803,Bit_1804,Bit_1805,Bit_1806,Bit_1807,Bit_1808,Bit_1809,Bit_1810,Bit_1811,Bit_1812,Bit_1813,Bit_1814,Bit_1815,Bit_1816,Bit_1817,Bit_1818,Bit_1819,Bit_1820,Bit_1821,Bit_1822,Bit_1823,Bit_1824,Bit_1825,Bit_1826,Bit_1827,Bit_1828,Bit_1829,Bit_1830,Bit_1831,Bit_1832,Bit_1833,Bit_1834,Bit_1835,Bit_1836,Bit_1837,Bit_1838,Bit_1839,Bit_1840,Bit_1841,Bit_1842,Bit_1843,Bit_1844,Bit_1845,Bit_1846,Bit_1847,Bit_1848,Bit_1849,Bit_1850,Bit_1851,Bit_1852,Bit_1853,Bit_1854,Bit_1855,Bit_1856,Bit_1857,Bit_1858,Bit_1859,Bit_1860,Bit_1861,Bit_1862,Bit_1863,Bit_1864,Bit_1865,Bit_1866,Bit_1867,Bit_1868,Bit_1869,Bit_1870,Bit_1871,Bit_1872,Bit_1873,Bit_1874,Bit_1875,Bit_1876,Bit_1877,Bit_1878,Bit_1879,Bit_1880,Bit_1881,Bit_1882,Bit_1883,Bit_1884,Bit_1885,Bit_1886,Bit_1887,Bit_1888,Bit_1889,Bit_1890,Bit_1891,Bit_1892,Bit_1893,Bit_1894,Bit_1895,Bit_1896,Bit_1897,Bit_1898,Bit_1899,Bit_1900,Bit_1901,Bit_1902,Bit_1903,Bit_1904,Bit_1905,Bit_1906,Bit_1907,Bit_1908,Bit_1909,Bit_1910,Bit_1911,Bit_1912,Bit_1913,Bit_1914,Bit_1915,Bit_1916,Bit_1917,Bit_1918,Bit_1919,Bit_1920,Bit_1921,Bit_1922,Bit_1923,Bit_1924,Bit_1925,Bit_1926,Bit_1927,Bit_1928,Bit_1929,Bit_1930,Bit_1931,Bit_1932,Bit_1933,Bit_1934,Bit_1935,Bit_1936,Bit_1937,Bit_1938,Bit_1939,Bit_1940,Bit_1941,Bit_1942,Bit_1943,Bit_1944,Bit_1945,Bit_1946,Bit_1947,Bit_1948,Bit_1949,Bit_1950,Bit_1951,Bit_1952,Bit_1953,Bit_1954,Bit_1955,Bit_1956,Bit_1957,Bit_1958,Bit_1959,Bit_1960,Bit_1961,Bit_1962,Bit_1963,Bit_1964,Bit_1965,Bit_1966,Bit_1967,Bit_1968,Bit_1969,Bit_1970,Bit_1971,Bit_1972,Bit_1973,Bit_1974,Bit_1975,Bit_1976,Bit_1977,Bit_1978,Bit_1979,Bit_1980,Bit_1981,Bit_1982,Bit_1983,Bit_1984,Bit_1985,Bit_1986,Bit_1987,Bit_1988,Bit_1989,Bit_1990,Bit_1991,Bit_1992,Bit_1993,Bit_1994,Bit_1995,Bit_1996,Bit_1997,Bit_1998,Bit_1999,Bit_2000,Bit_2001,Bit_2002,Bit_2003,Bit_2004,Bit_2005,Bit_2006,Bit_2007,Bit_2008,Bit_2009,Bit_2010,Bit_2011,Bit_2012,Bit_2013,Bit_2014,Bit_2015,Bit_2016,Bit_2017,Bit_2018,Bit_2019,Bit_2020,Bit_2021,Bit_2022,Bit_2023,Bit_2024,Bit_2025,Bit_2026,Bit_2027,Bit_2028,Bit_2029,Bit_2030,Bit_2031,Bit_2032,Bit_2033,Bit_2034,Bit_2035,Bit_2036,Bit_2037,Bit_2038,Bit_2039,Bit_2040,Bit_2041,Bit_2042,Bit_2043,Bit_2044,Bit_2045,Bit_2046,Bit_2047,Bit_2048
CHEMBL365134,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,1,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,1,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0
CHEMBL187579,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,1,0,0,0,0,0,0,0,0,0,0,0,0,0
5.3.2 予測モデルの構築
いよいよ、生成した説明変数(および目的変数)のデータを読み込み、予測モデルを作成する。論文に従いランダムフォレストを用いた。GridSearchによるハイパーパラメータ探索や、5フォールドクロスバリデーション、評価指標もできるかぎり論文を再現したかったので、githubのソースを一部流用している。また、githubのコードでは、scikit-learnのpermutation_test_scoreメソッドでY-randamiztionを行っていたため、これも流用した。
import argparse
import csv
import pandas as pd
import numpy as np
import gzip
import _pickle as cPickle
from sklearn import metrics
from sklearn.ensemble import RandomForestClassifier
from sklearn.feature_selection import VarianceThreshold
from sklearn.model_selection import train_test_split, cross_validate, GridSearchCV
from sklearn.model_selection import permutation_test_score, StratifiedKFold
def calc_metrics_derived_from_confusion_matrix(metrics_name, y_true, y_predict):
tn, fp, fn, tp = metrics.confusion_matrix(y_true, y_predict).ravel()
# PPV, precision
# TP / TP + FP
if metrics_name in ["PPV", "precision"]:
return tp / (tp + fp)
# NPV
# TN / TN + FN
if metrics_name in ["NPV"]:
return tn / (tn + fn)
# sensitivity, recall, TPR
# TP / TP + FN
if metrics_name in ["sensitivity", "recall", "TPR"]:
return tp / (tp + fn)
# specificity
# TN / TN + FP
if metrics_name in ["specificity"]:
return tn / (tn + fp)
def calc_metrics(metrics_name, y_true, y_predict):
if metrics_name == "accuracy":
return metrics.accuracy_score(y_true, y_predict)
if metrics_name == "ba":
return metrics.balanced_accuracy_score(y_true, y_predict)
if metrics_name == "roc_auc":
return metrics.roc_auc_score(y_true, y_predict)
if metrics_name == "kappa":
return metrics.cohen_kappa_score(y_true, y_predict)
if metrics_name == "mcc":
return metrics.matthews_corrcoef(y_true, y_predict)
if metrics_name == "precision":
return metrics.precision_score(y_true, y_predict)
if metrics_name == "recall":
return metrics.recall_score(y_true, y_predict)
def main():
parser = argparse.ArgumentParser()
parser.add_argument("-input", type=str, required=True)
parser.add_argument("-output_model", type=str, required=True)
parser.add_argument("-output_report", type=str, required=True)
args = parser.parse_args()
df = pd.read_csv(args.input, index_col=0)
print(df.shape)
y_train = df['outcome'].to_numpy()
print(y_train)
X_train = df.iloc[:, 1:]
print(y_train.shape)
print(X_train.shape)
# Number of trees in random forest
n_estimators = [100, 250, 500, 750, 1000]
max_features = ['auto', 'sqrt']
criterion = ['gini', 'entropy']
class_weight = [None,'balanced',
{0:.9, 1:.1}, {0:.8, 1:.2}, {0:.7, 1:.3}, {0:.6, 1:.4},
{0:.4, 1:.6}, {0:.3, 1:.7}, {0:.2, 1:.8}, {0:.1, 1:.9}]
random_state = [24]
# Create the random grid
param_grid = {'n_estimators': n_estimators,
'max_features': max_features,
'criterion': criterion,
'random_state': random_state,
'class_weight': class_weight}
# setup model building
rf = GridSearchCV(RandomForestClassifier(), param_grid, n_jobs=-1, cv=5, verbose=1)
rf.fit(X_train, y_train)
print()
print('Best params: %s' % rf.best_params_)
print('Score: %.2f' % rf.best_score_)
rf_best = RandomForestClassifier(**rf.best_params_, n_jobs=-1)
rf_best.fit(X_train, y_train)
#一旦できたモデルを保存
with gzip.GzipFile(args.output_model, 'w') as f:
cPickle.dump(rf_best, f)
with gzip.GzipFile(args.output_model, 'r') as f:
rf_best = cPickle.load(f)
# Params
pred = []
ad = []
pred_proba = []
index = []
cross_val = StratifiedKFold(n_splits=5)
# Do 5-fold loop
for train_index, test_index in cross_val.split(X_train, y_train):
fold_model = rf_best.fit(X_train.iloc[train_index], y_train[train_index])
fold_pred = rf_best.predict(X_train.iloc[test_index])
fold_ad = rf_best.predict_proba(X_train.iloc[test_index])
pred.append(fold_pred)
ad.append(fold_ad)
pred_proba.append(fold_ad[:, 1])
index.append(test_index)
threshold_ad = 0.70
# Prepare results to export
fold_index = np.concatenate(index)
fold_pred = np.concatenate(pred)
fold_ad = np.concatenate(ad)
fold_pred_proba = np.concatenate(pred_proba)
fold_ad = (np.amax(fold_ad, axis=1) >= threshold_ad).astype(str)
five_fold_morgan = pd.DataFrame({'Prediction': fold_pred, 'AD': fold_ad, 'Proba': fold_pred_proba}, index=list(fold_index))
five_fold_morgan.AD[five_fold_morgan.AD == 'False'] = np.nan
five_fold_morgan.AD[five_fold_morgan.AD == 'True'] = five_fold_morgan.Prediction
five_fold_morgan.sort_index(inplace=True)
five_fold_morgan['y_train'] = pd.DataFrame(y_train)
# morgan stats
all_datas = []
datas = []
datas.append(calc_metrics("accuracy", five_fold_morgan['y_train'], five_fold_morgan['Prediction']))
datas.append(calc_metrics("ba", five_fold_morgan['y_train'], five_fold_morgan['Prediction']))
datas.append(calc_metrics("roc_auc", five_fold_morgan['y_train'], five_fold_morgan['Proba']))
datas.append(calc_metrics("kappa", five_fold_morgan['y_train'], five_fold_morgan['Prediction']))
datas.append(calc_metrics("mcc", five_fold_morgan['y_train'], five_fold_morgan['Prediction']))
datas.append(calc_metrics("precision", five_fold_morgan['y_train'], five_fold_morgan['Prediction']))
datas.append(calc_metrics("recall", five_fold_morgan['y_train'], five_fold_morgan['Prediction']))
datas.append(calc_metrics_derived_from_confusion_matrix("sensitivity", five_fold_morgan['y_train'], five_fold_morgan['Prediction']))
datas.append(calc_metrics_derived_from_confusion_matrix("PPV", five_fold_morgan['y_train'], five_fold_morgan['Prediction']))
datas.append(calc_metrics_derived_from_confusion_matrix("specificity", five_fold_morgan['y_train'], five_fold_morgan['Prediction']))
datas.append(calc_metrics_derived_from_confusion_matrix("NPV", five_fold_morgan['y_train'], five_fold_morgan['Prediction']))
datas.append(1)
all_datas.append(datas)
# morgan AD stats
morgan_ad = five_fold_morgan.dropna(subset=['AD'])
morgan_ad['AD'] = morgan_ad['AD'].astype(int)
coverage_morgan_ad = len(morgan_ad['AD']) / len(five_fold_morgan['y_train'])
datas = []
datas.append(calc_metrics("accuracy", morgan_ad['y_train'], morgan_ad['AD']))
datas.append(calc_metrics("ba", morgan_ad['y_train'], morgan_ad['AD']))
datas.append(calc_metrics("roc_auc", morgan_ad['y_train'], morgan_ad['Proba']))
datas.append(calc_metrics("kappa", morgan_ad['y_train'], morgan_ad['AD']))
datas.append(calc_metrics("mcc", morgan_ad['y_train'], morgan_ad['AD']))
datas.append(calc_metrics("precision", morgan_ad['y_train'], morgan_ad['AD']))
datas.append(calc_metrics("recall", morgan_ad['y_train'], morgan_ad['AD']))
datas.append(calc_metrics_derived_from_confusion_matrix("sensitivity", morgan_ad['y_train'], morgan_ad['AD']))
datas.append(calc_metrics_derived_from_confusion_matrix("PPV", morgan_ad['y_train'], morgan_ad['AD']))
datas.append(calc_metrics_derived_from_confusion_matrix("specificity", morgan_ad['y_train'], morgan_ad['AD']))
datas.append(calc_metrics_derived_from_confusion_matrix("NPV", morgan_ad['y_train'], morgan_ad['AD']))
datas.append(coverage_morgan_ad)
all_datas.append(datas)
# print stats
print('\033[1m' + '5-fold External Cross Validation Statistical Characteristcs of QSAR models developed morgan' + '\n' + '\033[0m')
morgan_5f_stats = pd.DataFrame(all_datas, columns=["accuracy", "ba", "roc_auc", "kappa", "mcc", "precision", "recall",
"sensitivity", "PPV", "specificity", "NPV", "Coverage"])
morgan_5f_stats.to_csv(args.output_report)
print(morgan_5f_stats)
# Y ramdomaization
permutations = 20
score, permutation_scores, pvalue = permutation_test_score(rf_best, X_train, y_train,
cv=5, scoring='balanced_accuracy',
n_permutations=permutations,
n_jobs=-1,
verbose=1,
random_state=24)
print('True score = ', score.round(2),
'\nY-randomization = ', np.mean(permutation_scores).round(2),
'\np-value = ', pvalue.round(4))
if __name__ == "__main__":
main()
これを実行するとoutput_modelで指定したパスに予測モデル本体が保存され、output_reportで指定したパスに構築された予測モデルの評価指標が出力される。その結果は次の通りとなった。
accuracy balanced_accuracy roc_auc kappa mcc precision recall sensitivity PPV specificity NPV Coverage
0.801980 0.713126 0.807507 0.490156 0.551662 0.9375 0.441176 0.441176 0.9375 0.985075 0.776471 1.000000
0.852941 0.736842 0.812030 0.564661 0.627215 1.0000 0.473684 0.473684 1.0000 1.000000 0.830508 0.673267
True score = 0.71
Y-randomization = 0.48
p-value = 0.0476
前半の1行目のスコアは、全学習データにおけるクロスバリデーションのスコアである。recallとsensitivityは同じことだが、scikit-learn版と自前で組み込んだ関数が一致しているかの比較のため、両方表示している。
前半の2行目は、論文の中で考慮されているものであり、予測信頼度(予測したクラスに対するscikit-learnのpredict_probaで得られる値といえばわかるだろうか)がしきい値(0.7)以上のデータに限定したクロスバリデーションのスコアであり、より信頼度の高いデータだけによる指標と考えられ、多少指標がよくなっている。
後半は、Y-randamizationの結果であり、scikit-learnのドキュメントによると、True scoreはターゲットを入れ替えない真のスコア、Y-randamizationはターゲットをランダマイズしたスコア、 p-valueはスコアが偶然に得られる確率であり、最良は1 /(n_permutations + 1)で最悪は1.0らしい。今回、1 /(n_permutations + 1)= 1 / (20 + 1) = 0.0467であり、最良の値になっているため、0.71 というbalanced_accuracyは偶然に得られたものではなさそうだ。
5.4 予測モデルを利用した候補薬物のスクリーニング
5.4.1 DrugBankデータに対する予測の実行
5.3で作成した予測モデル本体と、ChEMBL、PDBから抽出した学習データ(101件)、DrugBankから抽出したバーチャルスクリーニング用データ(10750件)をそれぞれ以下のスクリプト(断片のみ記載)に指定して、予測を行う。
parser = argparse.ArgumentParser()
parser.add_argument("-input_train", type=str, required=True)
parser.add_argument("-input_predict", type=str, required=True)
parser.add_argument("-input_model", type=str, required=True)
parser.add_argument("-output_csv", type=str, required=True)
parser.add_argument("-output_report", type=str, required=True)
args = parser.parse_args()
#学習データ読み込み
trains = list()
with open(args.input_train, "r") as f:
reader = csv.DictReader(f)
for i, row in enumerate(reader):
smiles = row["canonical_smiles"]
new_smiles = normalize(smiles)
trains.append((row["id"], new_smiles))
#予測データの読み込み
smiles_list = list()
datas = []
datas_other = []
with open(args.input_predict, "r") as f:
reader = csv.DictReader(f)
for row in reader:
print(row["id"])
smiles = row["smiles"]
new_smiles = normalize(smiles)
dup_pred = None
dup_train = None
if new_smiles is None or Chem.MolFromSmiles(new_smiles) is None:
print(f"error! {id}")
new_smiles = smiles
existFlag = False
for db_id, tmp_smiles in smiles_list:
if new_smiles == tmp_smiles:
dup_pred = db_id
print(f"{id}: same structure exist in predict data! - {db_id}, {tmp_smiles}")
break
for db_id, tmp_smiles in trains:
if new_smiles == tmp_smiles:
dup_train = db_id
print(f"{id}: same structure exist in train data! - {db_id}, {tmp_smiles}")
break
smiles_list.append((row["id"], new_smiles))
mol = Chem.MolFromSmiles(new_smiles)
fp = AllChem.GetMorganFingerprintAsBitVect(mol, radius=3, nBits=2048, useFeatures=False, useChirality=False)
fp = pd.Series(np.asarray(fp)).values
data = []
data.append(row["id"])
data.extend(fp)
datas.append(data)
datas_other.append((row["id"], row["name"], row["status"], dup_pred, dup_train))
columns = list()
columns.append("id")
columns.extend(["Bit_" + str(i+1) for i in range(2048)])
df = pd.DataFrame(data=datas, columns=columns)
df.set_index("id", inplace=True, drop=True)
df_other = pd.DataFrame(data=datas_other, columns=["id", "name", "status", "dup_predict", "dup_train"])
df_other.set_index("id", inplace=True, drop=True)
#一旦保存
df.to_csv(args.output_csv)
#df = pd.read_csv(args.input, index_col=0)
X_vs = df.iloc[:, 0:]
with gzip.GzipFile(args.input_model, 'r') as f:
model = cPickle.load(f)
ad_threshold = 0.70
y_pred = model.predict(X_vs)
confidence = model.predict_proba(X_vs)
confidence = np.amax(confidence, axis=1).round(2)
ad = confidence >= ad_threshold
pred = pd.DataFrame({'Prediction': y_pred, 'AD': ad, 'Confidence': confidence}, index=None)
pred.AD[pred.AD == False] = np.nan
pred.AD[pred.AD == True] = pred.Prediction.astype(int)
pred_ad = pred.dropna().astype(int)
coverage_ad = len(pred_ad) * 100 / len(pred)
print('VS pred: %s' % pred.Prediction)
print('VS pred AD: %s' % pred_ad.Prediction)
print('Coverage of AD: %.2f%%' % coverage_ad)
pred.index = X_vs.index
predictions = pd.concat([df_other, pred], axis=1)
for col in ['Prediction', 'AD']:
predictions[col].replace(0, 'Inactive', inplace=True)
predictions[col].replace(1, 'Active', inplace=True)
print(predictions.head())
predictions.to_csv(args.output_report)
output_csvには予測データの説明変数の計算結果が出力される。また、output_reportにはDrugBankの各薬剤のDrugBank ID, Common Name, 市販/中止/実験等の種別、予測結果、しきい値を超える予測信頼度に対する予測結果、予測信頼度等が出力される。また、outptu_reportには、各予測データにおいて、同じ予測データの中で重複する構造があった場合および学習データに対して重複する構造があった場合、それぞれ重複したIDを出力するようにした。予測データの中で321件、学習データに対して6件、重複するデータ存在した。
5.4.2 論文との比較
論文では、DrugBankに対するバーチャルスクリーニング結果は以下に記載されている。
https://github.com/alvesvm/sars-cov-mpro/tree/master/mol-inf-2020/datasets/screened_compounds
drugbank_hits_qsar_consensus.xlsx
ここに出力されているDrugBankの薬剤およびその結果と、前項で出力した予測結果を比較した。
論文では、SiRIMS、Dragon, そして今回作成したRDKitのMorganの3パターンの記述子計算による予測モデルを作成し、そのうち複数のモデルがActiveとしたものを最終的なActiveとしている。
全体的な比較と、SiRIMS、Dragonそれぞれに対する比較をしてみた。
なお、github上のMorganの結果は気になる点を見つけたため、これとの比較は見送っている。
全体的な結果との比較
論文では41件となっていたが、github上のエクセルでは最終的なActive件数は51件となっていた。
その51件に対し、今回作成したモデルがActiveとしたものは、19件であった。
SiRIMSモデル、Dragonモデルとの比較
SiRIMSモデル、Dragonモデルと、Active/Inactive件数の比較は以下の通りである。
| MODEL | ACTIVE | INACITVE | TOTAL |
|---|---|---|---|
| SiRMS | 309 | 9305 | 9614 |
| DRAGON | 864 | 8750 | 9614 |
| 作成モデル | 314 | 9300 | 9614 |
つづいてSiRMISモデル、DRragonモデルそれぞれと共通のActive数を比較してみた。
| MODEL | SiRMS | DRAGON | 作成モデル |
|---|---|---|---|
| SiRMS | - | 39 | 79 |
| DRAGON | 39 | - | 35 |
| 作成モデル | 79 | 35 | - |
Active数はDRAGONが最も多いにもかかわらず、今回の作成したモデルと一致したActive件数は、SiRMSが79件でありDragonの倍になっている。このことから今回の作成モデルはDragonよりはSiRMSに類似したモデルと考えられる。
5.4.3 おまけ: 予測モデルの適用範囲の可視化
論文には記載がなかったものの、学習データ上に予測対象のデータがどのように分布するかを可視化してみた。機械学習では学習データと似ていないデータの場合、予測の信頼性が問題となるからである。方法としてはまず、学習データで次元削減モデルを作成し、そのモデルを用いて予測対象データの2次元座標を生成しプロットしてみた。
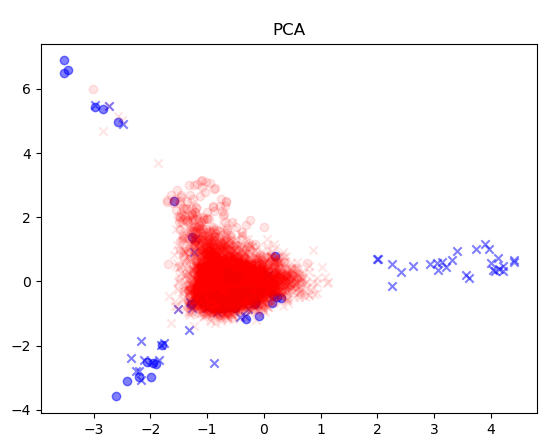
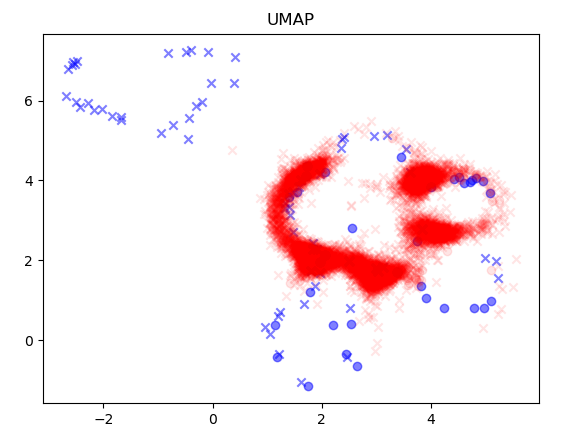
次元削減手法はPCAとUMAPを試した。データは作成モデルと同様、Morganフィンガープリントを利用した。
青が学習データ、赤が予測データである。〇がActive、XがInactiveになる。学習データで〇とXが重なっているところは、UMAPの方がうまく分離できているように見える。
可視化したスクリプトのソースは以下の通りとなる。学習データでPCAまたはUMAPのモデルをfitで作成し、それに対し各予測データをtransformした結果を表示している。図をみると予測対象データは、学習データの範囲から大きく逸脱していないように見えるが、学習データから構成されるデータのみで次元削減していることも影響しているかもしれない。
parser = argparse.ArgumentParser()
parser.add_argument("-train", type=str, required=True)
parser.add_argument("-predict", type=str)
parser.add_argument("-result", type=str)
parser.add_argument("-method", type=str, default="PCA", choices=["PCA", "UMAP"])
args = parser.parse_args()
# all_train.csvの読み込み, fpの計算
train_datas = []
train_datas_active = []
train_datas_inactive = []
with open(args.train, "r") as f:
reader = csv.DictReader(f)
for row in reader:
smiles = row["canonical_smiles"]
mol = Chem.MolFromSmiles(smiles)
fp = AllChem.GetMorganFingerprintAsBitVect(mol, radius=3, nBits=2048, useFeatures=False, useChirality=False)
train_datas.append(fp)
if int(row["outcome"]) == 1:
train_datas_active.append(fp)
else:
train_datas_inactive.append(fp)
if args.predict and args.result:
result_outcomes = []
result_ads = []
# 予測結果読み込み
with open(args.result, "r",encoding="utf-8_sig") as f:
reader = csv.DictReader(f)
for i, row in enumerate(reader):
#print(row)
if row["Prediction"] == "Active":
result_outcomes.append(1)
else:
result_outcomes.append(0)
result_ads.append(row["Confidence"])
# drugbank.csvの読み込み, fpの計算
predict_datas = []
predict_datas_active = []
predict_datas_inactive = []
predict_ads = []
with open(args.predict, "r") as f:
reader = csv.DictReader(f)
for i, row in enumerate(reader):
print(i)
smiles = row["smiles"]
mol = Chem.MolFromSmiles(smiles)
fp = AllChem.GetMorganFingerprintAsBitVect(mol, radius=3, nBits=2048, useFeatures=False, useChirality=False)
predict_datas.append(fp)
if result_outcomes[i] == 1:
predict_datas_active.append(fp)
else:
predict_datas_inactive.append(fp)
# 分析
model = None
if args.method == "PCA":
model = PCA(n_components=2)
model.fit(train_datas)
if args.method == "UMAP":
model = umap.UMAP()
model.fit(train_datas)
result_train = model.transform(train_datas)
result_train_active = model.transform(train_datas_active)
result_train_inactive = model.transform(train_datas_inactive)
plt.title(args.method)
#plt.scatter(result_train[:, 0], result_train[:, 1], c="blue", alpha=0.1, marker="o")
plt.scatter(result_train_active[:, 0], result_train_active[:, 1], c="blue", alpha=0.5, marker="o")
plt.scatter(result_train_inactive[:, 0], result_train_inactive[:, 1], c="blue", alpha=0.5, marker="x")
# 予測(predict)
if args.predict and args.result:
result_predict = model.transform(predict_datas)
result_predict_active = model.transform(predict_datas_active)
result_predict_inactive = model.transform(predict_datas_inactive)
#plt.scatter(result_predict[:, 0], result_predict[:, 1], c=result_ads, alpha=0.1, cmap='viridis_r')
plt.scatter(result_predict_active[:, 0], result_predict_active[:, 1], c="red", alpha=0.1, marker="o")
plt.scatter(result_predict_inactive[:, 0], result_predict_inactive[:, 1], c="red", alpha=0.1, marker="x")
plt.show()
6. おわりに
6.1 感想
今回、論文の内容を試した過程を通じて、以下のことを得ることができた。今後自ら解析を行ったり、別の論文を読んだりする場合に多いに役立つと思う。
- 創薬分野におけるデータ収集から前処理、予測モデル作成、予測モデルの評価までを一通り体験することができた。特に予測モデルを作成する前のデータの収集や前処理、予測モデルを作成した後の適用・評価がいかに手間がかかるのかか身をもって体験することができた。
- 公共データべースについて、概要やデータ収集方法などの理解を深めることができた。
今回は従来手法による論文を試したが、Graph Convolution Networks(GCN) などDeep Learning を活用した論文も続々と出てきており、近いうちにそれらも試してみたい。
6.2 今回の論文に関連したその他のお勧めの論文
論文を探す過程で、他に見つかったコロナウィルス関連の予測モデルに関する論文を以下に2点あげる。いずれも、データセットが公開されているため試してみるにはよいと思う。
この論文では、本記事の論文とは異なるpapain-like protease (PLpro)いうターゲットに対する予測モデルを作成している。
これは、本記事の論文と同じメインプロテアーゼに対する予測モデルを作成している。OECD ガイドライン(https://www.oecd.org/chemicalsafety/risk-assessment/37849783.pdf )に沿ってモデルが評価されており、化合物を対象とした予測モデルの評価に関する知見を得ることができそうである。
6.3 本記事で利用したライブラリ
本記事で利用した主なライブラリは以下のとおりである。
- Python 3.7
- RDkit 2020.03.5
- scikit-learn 0.23.2
- MolVS 0.1.1
6.5 本記事のプログラムについて
本記事に掲載したプログラムは https://github.com/kimisyo/sars_trace_1 にて公開している。