はじめに
昨年の12月にscikit-learnの0.24.0がリリースされました。
https://scikit-learn.org/stable/whats_new/v0.24.html#version-0-24-0
partial_dependence の機能が拡張されたので、試してみました。
環境
環境は以下の通り。
$sw_vers
ProductName: Mac OS X
ProductVersion: 10.13.6
BuildVersion: 17G14042
Jupyterlab (Version 0.35.4) 上で作業していたので、python kernelのバージョンも記載しておきます。
Python 3.7.3 (default, Mar 27 2019, 16:54:48)
IPython 7.4.0 -- An enhanced Interactive Python. Type '?' for help.
やったこと
scikit-learnのバージョンアップ
Anacondaの環境を使っているので、condaでバージョンアップします。
conda install -c conda-forge scikit-learn=0.24.0
scikit-learnとこのあと使うlightgbmのバージョンは次の通りです。
from sklearn import __version__ as sk_ver
print(lgb.__version__)
print(sk_ver)
3.1.1
0.24.0
モデル構築
モデルを用意します。
データはscikit-learnで用意されているボストンデータセットを使用しました。
import pandas as pd
import sklearn.datasets as skd
data = skd.load_boston()
df_X = pd.DataFrame(data.data, columns=data.feature_names)
df_y = pd.DataFrame(data.target, columns=['y'])
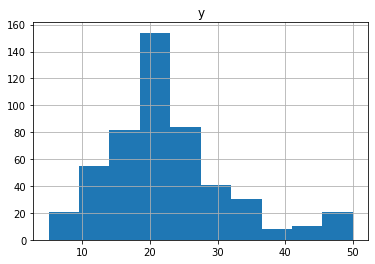
目的変数の分布は下図の通りで、20あたりにピークを持つような形になっています。
説明変数については、506行13列のデータで全列がnon-nullのfloat型なのでこのままモデルを作ります。
以前の記事で述べましたが、APIの仕様を合わせるため、LightgbmはScikit-learn APIで学習させます。
import lightgbm as lgb
from sklearn.model_selection import train_test_split
df_X_train, df_X_test, df_y_train, df_y_test = train_test_split(df_X, df_y, test_size=0.2, random_state=4)
lgbm_sk = lgb.LGBMRegressor(objective='regression',
random_state=4,
metric='rmse')
lgbm_sk.fit(df_X_train, df_y_train)
LGBMRegressor(metric='rmse', objective='regression', random_state=4)
partial-dependenceの変更箇所と試したこと
partial_dependence のドキュメントを見ると、引数は次のようになっています。
sklearn.inspection.partial_dependence(estimator, X, features, *, response_method='auto', percentiles=0.05, 0.95, grid_resolution=100, method='auto', kind='legacy')
このうち、kindのパラメータが0.24.0で追加されました。
kindについてドキュメントでは
kind{‘legacy’, ‘average’, ‘individual’, ‘both’}, default=’legacy’
Whether to return the partial dependence averaged across all the samples in the dataset or one line per sample or both. See Returns below.Note that the fast method='recursion' option is only available for kind='average'. Plotting individual dependencies requires using the slower method='brute' option.
New in version 0.24.
Deprecated since version 0.24: kind='legacy' is deprecated and will be removed in version 1.1. kind='average' will be the new default. It is intended to migrate from the ndarray output to Bunch output.
と記載されています。現デフォルトのlegacyはそのうちなくなるとのことですが、‘legacy’, ‘average’, ‘individual’, ‘both’の4種類を試してみました。
kind = 'legacy'
kind = 'legacy' でCRIMという説明変数についてのpartial dependenceを出してみます。
from sklearn.inspection import partial_dependence
result_pd_legacy = partial_dependence(lgbm_sk, features=['CRIM'], percentiles=(0,1), X=df_X_train, kind='legacy')
result_pd_legacy
返り値はndarrayとListがtupleで入っているような形になっています。
(array([[22.34455464, 22.80833098, 22.86597834, 22.96513457, 23.27716791,
23.47015226, 23.53235471, 23.35070292, 23.20272559, 23.07065778,
23.1786901 , 22.87241557, 22.59017997, 22.54737655, 22.54737655,
22.31672465, 22.31672465, 22.31672465, 22.18968352, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113]]),
[array([6.32000000e-03, 9.05005657e-01, 1.80369131e+00, 2.70237697e+00,
3.60106263e+00, 4.49974828e+00, 5.39843394e+00, 6.29711960e+00,
7.19580525e+00, 8.09449091e+00, 8.99317657e+00, 9.89186222e+00,
1.07905479e+01, 1.16892335e+01, 1.25879192e+01, 1.34866048e+01,
1.43852905e+01, 1.52839762e+01, 1.61826618e+01, 1.70813475e+01,
1.79800331e+01, 1.88787188e+01, 1.97774044e+01, 2.06760901e+01,
2.15747758e+01, 2.24734614e+01, 2.33721471e+01, 2.42708327e+01,
2.51695184e+01, 2.60682040e+01, 2.69668897e+01, 2.78655754e+01,
2.87642610e+01, 2.96629467e+01, 3.05616323e+01, 3.14603180e+01,
3.23590036e+01, 3.32576893e+01, 3.41563749e+01, 3.50550606e+01,
3.59537463e+01, 3.68524319e+01, 3.77511176e+01, 3.86498032e+01,
3.95484889e+01, 4.04471745e+01, 4.13458602e+01, 4.22445459e+01,
4.31432315e+01, 4.40419172e+01, 4.49406028e+01, 4.58392885e+01,
4.67379741e+01, 4.76366598e+01, 4.85353455e+01, 4.94340311e+01,
5.03327168e+01, 5.12314024e+01, 5.21300881e+01, 5.30287737e+01,
5.39274594e+01, 5.48261451e+01, 5.57248307e+01, 5.66235164e+01,
5.75222020e+01, 5.84208877e+01, 5.93195733e+01, 6.02182590e+01,
6.11169446e+01, 6.20156303e+01, 6.29143160e+01, 6.38130016e+01,
6.47116873e+01, 6.56103729e+01, 6.65090586e+01, 6.74077442e+01,
6.83064299e+01, 6.92051156e+01, 7.01038012e+01, 7.10024869e+01,
7.19011725e+01, 7.27998582e+01, 7.36985438e+01, 7.45972295e+01,
7.54959152e+01, 7.63946008e+01, 7.72932865e+01, 7.81919721e+01,
7.90906578e+01, 7.99893434e+01, 8.08880291e+01, 8.17867147e+01,
8.26854004e+01, 8.35840861e+01, 8.44827717e+01, 8.53814574e+01,
8.62801430e+01, 8.71788287e+01, 8.80775143e+01, 8.89762000e+01])])
2つ目のListはfeaturesのパラメータの入力に掃討していて、今はCRIMのみ指定しているので、中身は100個分(パラメータgridのデフォルト値)のCRIMのデータポイントになっています。
1つ目のndarrayの中身が、各データポイントに対する予測値の平均となっています。
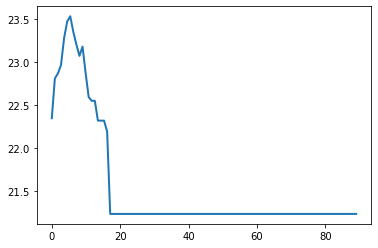
これを図示すると、いわゆるpartial dependenceのプロットを得られます。
plt.plot(result_pd_legacy[1][0].tolist(), result_pd_legacy[0][0].tolist(), lw=2)
plt.show()
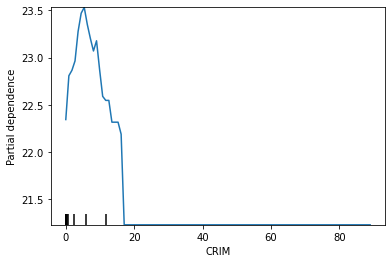
scikit-learnのplot_partial_dependenceでも図示してみると、上を同じことがわかります。
from sklearn.inspection import plot_partial_dependence
plot_partial_dependence(lgbm_sk, features=['CRIM'], percentiles=(0,1), X=df_X_train)
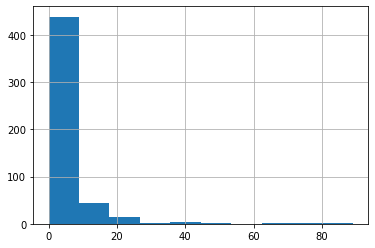
ちなみにCRIMの分布は下図の通りです。
df_X['CRIM'].hist()
kind = 'individual'
次に、kind = 'individual' でCRIMについてのpartial dependenceを出してみます。
result_pd_individual = partial_dependence(lgbm_sk, features=['CRIM'], percentiles=(0,1), X=df_X_train, kind='individual')
result_pd_individual
返り値はsklearn.utils.Bunch型になっています。
{'individual': array([[[35.65076711, 36.80808717, 36.80808717, ..., 36.30216375,
36.30216375, 36.30216375],
[13.75253977, 13.88194832, 13.88194832, ..., 11.88607485,
11.88607485, 11.88607485],
[25.40068607, 26.19198637, 26.19198637, ..., 26.32272304,
26.32272304, 26.32272304],
...,
[13.44052832, 13.37887547, 13.49894852, ..., 9.30373171,
9.30373171, 9.30373171],
[22.68308121, 24.31486889, 24.30242905, ..., 23.9277052 ,
23.9277052 , 23.9277052 ],
[20.91061255, 19.55812895, 19.54568912, ..., 16.7640575 ,
16.7640575 , 16.7640575 ]]]),
'values': [array([6.32000000e-03, 9.05005657e-01, 1.80369131e+00, 2.70237697e+00,
3.60106263e+00, 4.49974828e+00, 5.39843394e+00, 6.29711960e+00,
7.19580525e+00, 8.09449091e+00, 8.99317657e+00, 9.89186222e+00,
1.07905479e+01, 1.16892335e+01, 1.25879192e+01, 1.34866048e+01,
1.43852905e+01, 1.52839762e+01, 1.61826618e+01, 1.70813475e+01,
1.79800331e+01, 1.88787188e+01, 1.97774044e+01, 2.06760901e+01,
2.15747758e+01, 2.24734614e+01, 2.33721471e+01, 2.42708327e+01,
2.51695184e+01, 2.60682040e+01, 2.69668897e+01, 2.78655754e+01,
2.87642610e+01, 2.96629467e+01, 3.05616323e+01, 3.14603180e+01,
3.23590036e+01, 3.32576893e+01, 3.41563749e+01, 3.50550606e+01,
3.59537463e+01, 3.68524319e+01, 3.77511176e+01, 3.86498032e+01,
3.95484889e+01, 4.04471745e+01, 4.13458602e+01, 4.22445459e+01,
4.31432315e+01, 4.40419172e+01, 4.49406028e+01, 4.58392885e+01,
4.67379741e+01, 4.76366598e+01, 4.85353455e+01, 4.94340311e+01,
5.03327168e+01, 5.12314024e+01, 5.21300881e+01, 5.30287737e+01,
5.39274594e+01, 5.48261451e+01, 5.57248307e+01, 5.66235164e+01,
5.75222020e+01, 5.84208877e+01, 5.93195733e+01, 6.02182590e+01,
6.11169446e+01, 6.20156303e+01, 6.29143160e+01, 6.38130016e+01,
6.47116873e+01, 6.56103729e+01, 6.65090586e+01, 6.74077442e+01,
6.83064299e+01, 6.92051156e+01, 7.01038012e+01, 7.10024869e+01,
7.19011725e+01, 7.27998582e+01, 7.36985438e+01, 7.45972295e+01,
7.54959152e+01, 7.63946008e+01, 7.72932865e+01, 7.81919721e+01,
7.90906578e+01, 7.99893434e+01, 8.08880291e+01, 8.17867147e+01,
8.26854004e+01, 8.35840861e+01, 8.44827717e+01, 8.53814574e+01,
8.62801430e+01, 8.71788287e+01, 8.80775143e+01, 8.89762000e+01])]}
Bunchの中に'individual'と'values'というkeyがあってそれぞれndarrayとListになっています。
kind = 'legacy'のときの返り値に対応していますが、'individual'の出力がちょっと違います。ndarrayのshapeをみて見ると、
result_pd_individual['individual'].shape
(1, 404, 100)
という形になっています。404というのは、partial_dependenceに渡しているdf_X_trainの行数になります。
len(df_X_train)
404
ということで、いままでは予測値の平均しか返ってきていなかったですが、各データの予測値そのものが返るようになっています。
例えば、CRIMのgridの1点目に対して、404個の予測値の平均をとると、
result_pd_individual['individual'][0,:,0].mean()
22.344554644753032
となって、kind = 'legacy'で得られた1点目の予測値の平均22.34455464と一致します。
kind = 'average'
次に、kind = 'average' でCRIMについてのpartial dependenceを出してみます。
result_pd_average = partial_dependence(lgbm_sk, features=['CRIM'], percentiles=(0,1), X=df_X_train, kind='average')
result_pd_average
返り値はsklearn.utils.Bunch型になっています。
{'average': array([[22.34455464, 22.80833098, 22.86597834, 22.96513457, 23.27716791,
23.47015226, 23.53235471, 23.35070292, 23.20272559, 23.07065778,
23.1786901 , 22.87241557, 22.59017997, 22.54737655, 22.54737655,
22.31672465, 22.31672465, 22.31672465, 22.18968352, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113]]), 'values': [array([6.32000000e-03, 9.05005657e-01, 1.80369131e+00, 2.70237697e+00,
3.60106263e+00, 4.49974828e+00, 5.39843394e+00, 6.29711960e+00,
7.19580525e+00, 8.09449091e+00, 8.99317657e+00, 9.89186222e+00,
1.07905479e+01, 1.16892335e+01, 1.25879192e+01, 1.34866048e+01,
1.43852905e+01, 1.52839762e+01, 1.61826618e+01, 1.70813475e+01,
1.79800331e+01, 1.88787188e+01, 1.97774044e+01, 2.06760901e+01,
2.15747758e+01, 2.24734614e+01, 2.33721471e+01, 2.42708327e+01,
2.51695184e+01, 2.60682040e+01, 2.69668897e+01, 2.78655754e+01,
2.87642610e+01, 2.96629467e+01, 3.05616323e+01, 3.14603180e+01,
3.23590036e+01, 3.32576893e+01, 3.41563749e+01, 3.50550606e+01,
3.59537463e+01, 3.68524319e+01, 3.77511176e+01, 3.86498032e+01,
3.95484889e+01, 4.04471745e+01, 4.13458602e+01, 4.22445459e+01,
4.31432315e+01, 4.40419172e+01, 4.49406028e+01, 4.58392885e+01,
4.67379741e+01, 4.76366598e+01, 4.85353455e+01, 4.94340311e+01,
5.03327168e+01, 5.12314024e+01, 5.21300881e+01, 5.30287737e+01,
5.39274594e+01, 5.48261451e+01, 5.57248307e+01, 5.66235164e+01,
5.75222020e+01, 5.84208877e+01, 5.93195733e+01, 6.02182590e+01,
6.11169446e+01, 6.20156303e+01, 6.29143160e+01, 6.38130016e+01,
6.47116873e+01, 6.56103729e+01, 6.65090586e+01, 6.74077442e+01,
6.83064299e+01, 6.92051156e+01, 7.01038012e+01, 7.10024869e+01,
7.19011725e+01, 7.27998582e+01, 7.36985438e+01, 7.45972295e+01,
7.54959152e+01, 7.63946008e+01, 7.72932865e+01, 7.81919721e+01,
7.90906578e+01, 7.99893434e+01, 8.08880291e+01, 8.17867147e+01,
8.26854004e+01, 8.35840861e+01, 8.44827717e+01, 8.53814574e+01,
8.62801430e+01, 8.71788287e+01, 8.80775143e+01, 8.89762000e+01])]}
Bunchの中に'average'と'values'というkeyがあってそれぞれndarrayとListになっています。
'average'の中身はkind = 'legacy'のときの返り値と同じですので、前バージョンまでの出力に相当しています。
kind = 'both'
次に、kind = 'both' でCRIMについてのpartial dependenceを出してみます。
result_pd_both = partial_dependence(lgbm_sk, features=['CRIM'], percentiles=(0,1), X=df_X_train, kind='both')
result_pd_both
{'average': array([[22.34455464, 22.80833098, 22.86597834, 22.96513457, 23.27716791,
23.47015226, 23.53235471, 23.35070292, 23.20272559, 23.07065778,
23.1786901 , 22.87241557, 22.59017997, 22.54737655, 22.54737655,
22.31672465, 22.31672465, 22.31672465, 22.18968352, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113,
21.23030113, 21.23030113, 21.23030113, 21.23030113, 21.23030113]]), 'individual': array([[[35.65076711, 36.80808717, 36.80808717, ..., 36.30216375,
36.30216375, 36.30216375],
[13.75253977, 13.88194832, 13.88194832, ..., 11.88607485,
11.88607485, 11.88607485],
[25.40068607, 26.19198637, 26.19198637, ..., 26.32272304,
26.32272304, 26.32272304],
...,
[13.44052832, 13.37887547, 13.49894852, ..., 9.30373171,
9.30373171, 9.30373171],
[22.68308121, 24.31486889, 24.30242905, ..., 23.9277052 ,
23.9277052 , 23.9277052 ],
[20.91061255, 19.55812895, 19.54568912, ..., 16.7640575 ,
16.7640575 , 16.7640575 ]]]), 'values': [array([6.32000000e-03, 9.05005657e-01, 1.80369131e+00, 2.70237697e+00,
3.60106263e+00, 4.49974828e+00, 5.39843394e+00, 6.29711960e+00,
7.19580525e+00, 8.09449091e+00, 8.99317657e+00, 9.89186222e+00,
1.07905479e+01, 1.16892335e+01, 1.25879192e+01, 1.34866048e+01,
1.43852905e+01, 1.52839762e+01, 1.61826618e+01, 1.70813475e+01,
1.79800331e+01, 1.88787188e+01, 1.97774044e+01, 2.06760901e+01,
2.15747758e+01, 2.24734614e+01, 2.33721471e+01, 2.42708327e+01,
2.51695184e+01, 2.60682040e+01, 2.69668897e+01, 2.78655754e+01,
2.87642610e+01, 2.96629467e+01, 3.05616323e+01, 3.14603180e+01,
3.23590036e+01, 3.32576893e+01, 3.41563749e+01, 3.50550606e+01,
3.59537463e+01, 3.68524319e+01, 3.77511176e+01, 3.86498032e+01,
3.95484889e+01, 4.04471745e+01, 4.13458602e+01, 4.22445459e+01,
4.31432315e+01, 4.40419172e+01, 4.49406028e+01, 4.58392885e+01,
4.67379741e+01, 4.76366598e+01, 4.85353455e+01, 4.94340311e+01,
5.03327168e+01, 5.12314024e+01, 5.21300881e+01, 5.30287737e+01,
5.39274594e+01, 5.48261451e+01, 5.57248307e+01, 5.66235164e+01,
5.75222020e+01, 5.84208877e+01, 5.93195733e+01, 6.02182590e+01,
6.11169446e+01, 6.20156303e+01, 6.29143160e+01, 6.38130016e+01,
6.47116873e+01, 6.56103729e+01, 6.65090586e+01, 6.74077442e+01,
6.83064299e+01, 6.92051156e+01, 7.01038012e+01, 7.10024869e+01,
7.19011725e+01, 7.27998582e+01, 7.36985438e+01, 7.45972295e+01,
7.54959152e+01, 7.63946008e+01, 7.72932865e+01, 7.81919721e+01,
7.90906578e+01, 7.99893434e+01, 8.08880291e+01, 8.17867147e+01,
8.26854004e+01, 8.35840861e+01, 8.44827717e+01, 8.53814574e+01,
8.62801430e+01, 8.71788287e+01, 8.80775143e+01, 8.89762000e+01])]}
返り値はsklearn.utils.Bunch型になっていて、'average'と'individual'の両方が入っています。
自作のpartial dependenceプロット
せっかく各データ点の予測値が返るようになったので、ちょっと遊んでみました。
partial dependenceのプロットとして、平均値だけでなく、範囲と、とあるデータ点も一緒にプロットしてみます。
import matplotlib.pyplot as plt
import numpy as np
def pdp_plot(model, X, features, n_cols=3, pred_data=None):
n_cols = min(len(features), n_cols)
n_rows = int(np.ceil( len(features) / n_cols ))
fig = plt.figure(figsize=(4*n_cols, 4*n_rows))
ymin = 999
ymax = -999
for i, feature in enumerate(features):
result_pd_individual = partial_dependence(model,
features=[feature],
percentiles=(0.05,1-0.05),
X=X,
kind='individual')
feature_grid = result_pd_individual['values'][0]
pd_mean = result_pd_individual['individual'][0,:,:].mean(axis=0)
pd_perc_25pct = np.percentile(a=result_pd_individual['individual'][0,:,:], q=25, axis=0)
pd_perc_75pct = np.percentile(a=result_pd_individual['individual'][0,:,:], q=75, axis=0)
ax = fig.add_subplot(n_rows, n_cols, i+1)
ax.plot(feature_grid, pd_mean, label='average')
ax.fill_between(feature_grid, pd_perc_25pct, pd_perc_75pct, alpha=0.2, label='25 - 75 percentile')
if pred_data is not None:
pred = model.predict(pred_data)
ax.scatter(pred_data[feature], pred, marker='X', label='predicted data')
ax.set_xlabel(feature)
if (i+1) % n_cols == 1:
ax.set_ylabel('Partial dependence')
else:
ax.set_ylabel('')
ax.grid()
if i == 0:
ax.legend(loc=1)
ymin_, ymax_ = ax.get_ylim()
ymin = min(ymin_, ymin)
ymax = max(ymax_, ymax)
#plt.ylim(ymin*0.9, ymax*1.1)
for iax in fig.axes:
iax.set_ylim(ymin*0.9, ymax*1.1)
plt.show()
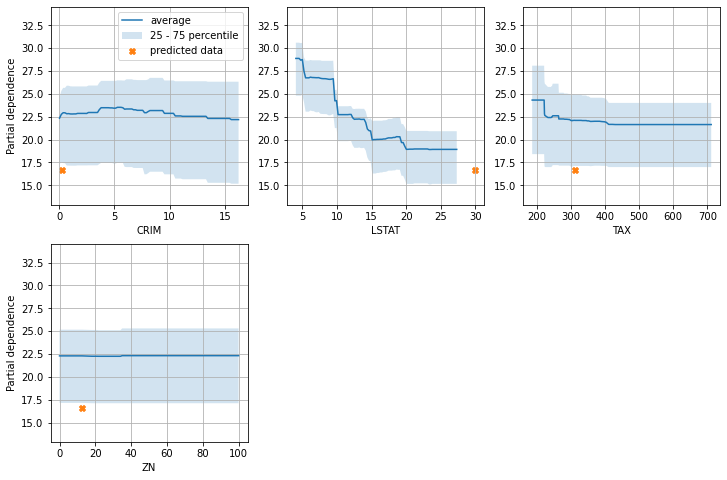
pdp_plot(model=lgbm_sk, X=df_X_train, features=['CRIM', 'LSTAT', 'TAX', 'ZN'], pred_data=pd.DataFrame([df_X_test.iloc[0, :]]))
とあるモデルの予測値の平均、25 - 75パーセンタイルの範囲の中に、とあるデータの説明変数の値+そのデータに対する予測値の情報を一緒にプロットしてみました。
なんとなくですが、なんでその予測値になったのかイメージつきやすいかなーと思った次第です。
まとめ
scikit-learnのバージョンが0.24.0になったことで、次の変更がありました。
- kindというパラメータが追加された。
- デフォルト('legacy')ではいままで通りの出力を得られる。ただし、今後'legacy'はなくなる。
- 今後のデフォルトは'average'となり、返り値はBunch型となる。
- 'individual'にすると各データに対する予測値を得られる。