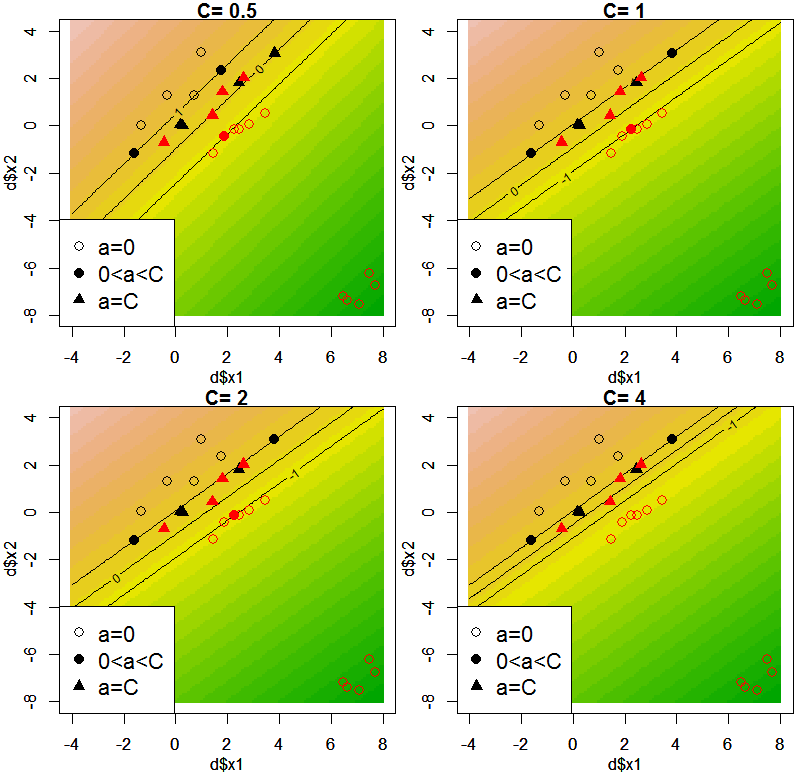
PRML 7.1.1に記載の、一部のデータの誤分類を許すようなサポートベクトルマシンを構成し、いくつかのCの値毎に、推定値、および各サンプルのラグランジュ乗数aを図示します。なお、カーネル関数は用いておらず、線形カーネル相当となります。二次計画法には、kernlabパッケージのipopを用いています。
library(MASS)
library(kernlab)
frame()
set.seed(0)
par(mfrow=c(2, 2))
par(mar=c(3, 3, 1, 0.1))
par(mgp=c(2, 1, 0))
xrange <- c(-4, 8)
yrange <- c(-8, 4)
x1 <- seq(xrange[1], xrange[2], .1)
x2 <- seq(yrange[1], yrange[2], .1)
x <- mvrnorm(10, c(0,1), matrix(c(2, 1, 1, 2), 2))
d1 <- as.data.frame(cbind(x, 1, 1))
x <- mvrnorm(10, c(2,0), matrix(c(2, 1, 1, 2), 2))
d1 <- rbind(d1, as.data.frame(cbind(x, 2, -1)))
x <- mvrnorm(5, c(7,-7), matrix(c(.2, 0, 0, .2), 2))
d2 <- rbind(d1, as.data.frame(cbind(x, 2, -1)))
d3 <- as.data.frame(rbind(cbind(1, 1, 1, 1), cbind(0, 1, 2, -1)))
names(d1) <- c("x1","x2","class","t")
names(d2) <- c("x1","x2","class","t")
names(d3) <- c("x1","x2","class","t")
doplot <- function(d, C) {
N <- nrow(d)
# quadratic programming
cc <- matrix(rep(-1, N))
hmat <- cbind(d$t * d$x1, d$t * d$x2)
hmat <- hmat %*% t(hmat)
ll <- matrix(rep(0, N))
uu <- matrix(rep(C, N))
bb <- 0
rr <- 0
amat <- matrix(d$t, nrow=1, ncol=N)
sv <- ipop(cc, hmat, amat, bb, ll, uu, rr)
a <- primal(sv)
# non-support vectors (a=0)
a[a < 1.0e-5] <- 0
# support vectors with penalty (a=C)
a[a > C - 1.0e-5] <- C
cat("a");print(a)
# derive b
bs <- c()
for (n in 1:N) {
if (a[n] > 0 && a[n] < C) {
bs <- c(bs, d$t[n] - rowSums(a * d$t * (cbind(d$x1[n], d$x2[n]) %*% t(cbind(d$x1, d$x2)))))
}
}
b <- mean(bs)
# plot estimates
estimate <- function(x1, x2) {
result <- 0
for (n in 1:N) {
if (a[n] != 0) {
result <- result + a[n] * d$t[n] * (cbind(x1, x2) %*% t(cbind(d$x1[n], d$x2[n])))
}
}
result <- result + b
}
y1 <- outer(x1, x2, estimate)
plot(d$x1, d$x2, col=d$class, type="n", xlim=xrange, ylim=yrange)
image(x1, x2, y1, xlim=xrange, ylim=yrange, col=terrain.colors(32), add=T)
contour(x1, x2, y1, xlim=xrange, ylim=yrange, levels=c(-1, 0, 1), add=T)
par(new=T)
plot(d$x1, d$x2, pch=ifelse(a == 0, 1, ifelse(a == C, 17, 16)), col=d$class, xlim=xrange, ylim=yrange, cex=1.3)
legend("bottomleft", legend=c("a=0","0<a<C","a=C"), pch=c(1, 16, 17), cex=1.3, y.intersp=1, bg="white")
title(paste("C=", C))
}
doplot(d2, 0.5)
doplot(d2, 1)
doplot(d2, 2)
doplot(d2, 4)