k-medoidsとは
暇な時に書きたい
k-medoidsのコード
import numpy as np
import sys
import random
from scipy.spatial.distance import cdist
from sklearn.preprocessing import LabelBinarizer
import matplotlib.pyplot as plt
class kMedoids(object):
#D_method is metric in distance.cdist
def __init__(self,n_cluster=4,max_iter=10,D_method="euclidean",init_method="random",eps=0):
self.n_cluster = n_cluster
self.max_iter = max_iter
self.D_method = D_method
self.D_matrix = None
self.labels = None
self.objective_value = float("inf")
self.loop_flag = True
self.eps = eps
self.lb = LabelBinarizer()
def calculate_D(self,data):
print("Calculate distance matrix (method is %s)"%self.D_method)
self.D_matrix = cdist(data,data,self.D_method)
return self.D_matrix
def _init_centroids(self):
l=list(range(self.D_matrix.shape[0]))
centroids = np.random.choice(l,n_cluster,replace=False)
return centroids
def _labels_to_membership(self):
if self.n_cluster == 2:
u = np.hstack((self.labels.reshape([self.n_data,1]),1-self.labels.reshape([self.n_data,1])))
else:
self.lb.fit(self.labels)
u = self.lb.transform(self.labels)
return u
def fit(self,data=None,D_matrix=None):
if D_matrix is None and data is None:
sys.stderr.write('Please give data or D_matrix as argument!!!')
return
elif data is not None:
self.D_matrix = self.calculate_D(data)
elif self.D_matrix is None:
self.D_matrix = self.calculate_D(data)
self.n_data = self.D_matrix.shape[0]
print("fit by k-Medoids")
centroids = self._init_centroids()
u = self._updateMembership(centroids)
num_loop = 0
while num_loop < self.max_iter and self.loop_flag:
centroids = self._updateCentroids(u)
u = self._updateMembership(centroids)
print("loop:%s objective_value:%s"%(num_loop,self.objective_value))
num_loop += 1
print("kMedoids is finish!!!")
self.loop_flag = True
def predict(self):
return self.labels
def _updateCentroids(self,u):
data_in_clusters = u==1
D_in_clusters = (data_in_clusters[np.newaxis,:,:]*data_in_clusters[:,np.newaxis,:]).swapaxes(0,2)
D_in_clusters = D_in_clusters*self.D_matrix[np.newaxis,:,:]
temp = np.sum(D_in_clusters,axis=2)
np.place(temp,temp==0,float("inf"))
centroids = np.argmin(temp,axis=1)
objective_value = np.sum(D_in_clusters[:,centroids])
if np.absolute(objective_value - self.objective_value) <= self.eps:
self.loop_flag = False
else:
self.objective_value = objective_value
return centroids
def _updateMembership(self,centroids):
self.labels = np.argmin(self.D_matrix[:,centroids],axis=1)
#print("test",self.labels)
u = self._labels_to_membership()
#print("test1",u)
return u
def objective_value(self):
return self.objective_value
テストデータ作成
from sklearn.datasets import make_blobs
data, a = make_blobs(n_samples=100,
n_features=2,
centers=3,
cluster_std=3.5,
shuffle=True
)
テストデータに対してkMedoidsを適用
n_cluster=4
km =kMedoids(n_cluster)
km.fit(data)
Calculate distance matrix (method is euclidean)
fit by k-Medoids
loop:0 objective_value:192.74729208157845
loop:1 objective_value:184.15560018589713
loop:2 objective_value:179.3414992176697
loop:3 objective_value:164.5655157771438
loop:4 objective_value:163.04503531227357
loop:5 objective_value:161.1913508723207
loop:6 objective_value:161.1913508723207
kMedoids is finish!!!
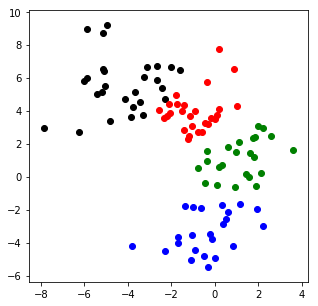
可視化
今後の課題
初期クラスタ中心の選択方法としてkmeas++的なのを追加する
実装が合ってるかはほとんどテストしてないのでテストすべき