tldr
KggleのAV : Healthcare Analytics II
をPredicting Hospital Stays - Data Every Day #024に沿ってやっていきます。
実行環境はGoogle Colaboratorです。
インポート
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import sklearn.preprocessing as sp
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
import tensorflow as tf
データのダウンロード
Google Driveをマウントします。
from google.colab import drive
drive.mount('/content/drive')
Mounted at /content/drive
KaggleのAPIクライアントを初期化し、認証します。
認証情報はGoogle Drive内(/content/drive/My Drive/Colab Notebooks/Kaggle)にkaggle.jsonとして置いてあります。
import os
kaggle_path = "/content/drive/My Drive/Colab Notebooks/Kaggle"
os.environ['KAGGLE_CONFIG_DIR'] = kaggle_path
from kaggle.api.kaggle_api_extended import KaggleApi
api = KaggleApi()
api.authenticate()
Kaggle APIを使ってデータをダウンロードします。
dataset_id = 'nehaprabhavalkar/av-healthcare-analytics-ii'
dataset = api.dataset_list_files(dataset_id)
file_name = dataset.files[0].name
file_path = os.path.join(api.get_default_download_dir(), file_name)
file_path
Warning: Looks like you're using an outdated API Version, please consider updating (server 1.5.10 / client 1.5.9)
'/content/healthcare/train_data.csv'
api.dataset_download_file(dataset_id, file_name, force=True, quiet=False)
100%|██████████| 4.45M/4.45M [00:00<00:00, 213MB/s]
Downloading train_data.csv.zip to /content
True
os.listdir('/content')
['.config', 'train_data.csv.zip', 'drive', 'sample_data']
import zipfile
zip_path = '/content/train_data.csv.zip'
with zipfile.ZipFile(zip_path) as existing_zip:
existing_zip.extractall('/content')
os.listdir('/content')
['.config', 'train_data.csv.zip', 'drive', 'train_data.csv', 'sample_data']
データの読み込み
Padasを使ってダウンロードしてきたCSVファイルを読み込みます。
data = pd.read_csv('/content/train_data.csv')
data
| case_id | Hospital_code | Hospital_type_code | City_Code_Hospital | Hospital_region_code | Available Extra Rooms in Hospital | Department | Ward_Type | Ward_Facility_Code | Bed Grade | patientid | City_Code_Patient | Type of Admission | Severity of Illness | Visitors with Patient | Age | Admission_Deposit | Stay | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 1 | 8 | c | 3 | Z | 3 | radiotherapy | R | F | 2.0 | 31397 | 7.0 | Emergency | Extreme | 2 | 51-60 | 4911.0 | 0-10 |
| 1 | 2 | 2 | c | 5 | Z | 2 | radiotherapy | S | F | 2.0 | 31397 | 7.0 | Trauma | Extreme | 2 | 51-60 | 5954.0 | 41-50 |
| 2 | 3 | 10 | e | 1 | X | 2 | anesthesia | S | E | 2.0 | 31397 | 7.0 | Trauma | Extreme | 2 | 51-60 | 4745.0 | 31-40 |
| 3 | 4 | 26 | b | 2 | Y | 2 | radiotherapy | R | D | 2.0 | 31397 | 7.0 | Trauma | Extreme | 2 | 51-60 | 7272.0 | 41-50 |
| 4 | 5 | 26 | b | 2 | Y | 2 | radiotherapy | S | D | 2.0 | 31397 | 7.0 | Trauma | Extreme | 2 | 51-60 | 5558.0 | 41-50 |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 318433 | 318434 | 6 | a | 6 | X | 3 | radiotherapy | Q | F | 4.0 | 86499 | 23.0 | Emergency | Moderate | 3 | 41-50 | 4144.0 | 11-20 |
| 318434 | 318435 | 24 | a | 1 | X | 2 | anesthesia | Q | E | 4.0 | 325 | 8.0 | Urgent | Moderate | 4 | 81-90 | 6699.0 | 31-40 |
| 318435 | 318436 | 7 | a | 4 | X | 3 | gynecology | R | F | 4.0 | 125235 | 10.0 | Emergency | Minor | 3 | 71-80 | 4235.0 | 11-20 |
| 318436 | 318437 | 11 | b | 2 | Y | 3 | anesthesia | Q | D | 3.0 | 91081 | 8.0 | Trauma | Minor | 5 | 11-20 | 3761.0 | 11-20 |
| 318437 | 318438 | 19 | a | 7 | Y | 5 | gynecology | Q | C | 2.0 | 21641 | 8.0 | Emergency | Minor | 2 | 11-20 | 4752.0 | 0-10 |
318438 rows × 18 columns
下準備
欠損値の処理
data.isnull().sum()
case_id 0
Hospital_code 0
Hospital_type_code 0
City_Code_Hospital 0
Hospital_region_code 0
Available Extra Rooms in Hospital 0
Department 0
Ward_Type 0
Ward_Facility_Code 0
Bed Grade 113
patientid 0
City_Code_Patient 4532
Type of Admission 0
Severity of Illness 0
Visitors with Patient 0
Age 0
Admission_Deposit 0
Stay 0
dtype: int64
欠損値に列全体の平均を挿入します。
def impute_missing_values(data, columns):
for column in columns:
data[column] = data[column].fillna(data[column].mean())
impute_columns = ['Bed Grade', 'City_Code_Patient']
impute_missing_values(data, impute_columns)
data.isnull().sum()
case_id 0
Hospital_code 0
Hospital_type_code 0
City_Code_Hospital 0
Hospital_region_code 0
Available Extra Rooms in Hospital 0
Department 0
Ward_Type 0
Ward_Facility_Code 0
Bed Grade 0
patientid 0
City_Code_Patient 0
Type of Admission 0
Severity of Illness 0
Visitors with Patient 0
Age 0
Admission_Deposit 0
Stay 0
dtype: int64
オブジェクト型の処理
data.dtypes
case_id int64
Hospital_code int64
Hospital_type_code object
City_Code_Hospital int64
Hospital_region_code object
Available Extra Rooms in Hospital int64
Department object
Ward_Type object
Ward_Facility_Code object
Bed Grade float64
patientid int64
City_Code_Patient float64
Type of Admission object
Severity of Illness object
Visitors with Patient int64
Age object
Admission_Deposit float64
Stay object
dtype: object
def get_categorical_unique(data):
# object型の列のみをリストにする
categorical_columns = [column for column in data.dtypes.index if data.dtypes[column] == 'object']
# 各列の値を取り出しマップを作る
categorical_uniques = {column: data[column].unique() for column in categorical_columns}
return categorical_uniques
get_categorical_unique(data)
{'Age': array(['51-60', '71-80', '31-40', '41-50', '81-90', '61-70', '21-30',
'11-20', '0-10', '91-100'], dtype=object),
'Department': array(['radiotherapy', 'anesthesia', 'gynecology', 'TB & Chest disease',
'surgery'], dtype=object),
'Hospital_region_code': array(['Z', 'X', 'Y'], dtype=object),
'Hospital_type_code': array(['c', 'e', 'b', 'a', 'f', 'd', 'g'], dtype=object),
'Severity of Illness': array(['Extreme', 'Moderate', 'Minor'], dtype=object),
'Stay': array(['0-10', '41-50', '31-40', '11-20', '51-60', '21-30', '71-80',
'More than 100 Days', '81-90', '61-70', '91-100'], dtype=object),
'Type of Admission': array(['Emergency', 'Trauma', 'Urgent'], dtype=object),
'Ward_Facility_Code': array(['F', 'E', 'D', 'B', 'A', 'C'], dtype=object),
'Ward_Type': array(['R', 'S', 'Q', 'P', 'T', 'U'], dtype=object)}
各データの特性を見て
- Labelエンコード
- Onehotエンコード
- Ordinalエンコード
のどれでエンコードするか判断する
Onehotエンコード
def onehot_encode(data, columns):
for column in columns:
dummies = pd.get_dummies(data[column])
data = pd.concat([data, dummies], axis=1)
data = data.drop(column, axis=1)
return data
onehot_columns = ['Department', 'Hospital_region_code', 'Hospital_type_code', 'Ward_Facility_Code', 'Ward_Type']
data = onehot_encode(data, onehot_columns)
Ordinalエンコード
categorical_uniques = get_categorical_unique(data)
categorical_uniques
{'Age': array(['51-60', '71-80', '31-40', '41-50', '81-90', '61-70', '21-30',
'11-20', '0-10', '91-100'], dtype=object),
'Severity of Illness': array(['Extreme', 'Moderate', 'Minor'], dtype=object),
'Stay': array(['0-10', '41-50', '31-40', '11-20', '51-60', '21-30', '71-80',
'More than 100 Days', '81-90', '61-70', '91-100'], dtype=object),
'Type of Admission': array(['Emergency', 'Trauma', 'Urgent'], dtype=object)}
for column in categorical_uniques:
categorical_uniques[column] = sorted(categorical_uniques[column])
categorical_uniques
{'Age': ['0-10',
'11-20',
'21-30',
'31-40',
'41-50',
'51-60',
'61-70',
'71-80',
'81-90',
'91-100'],
'Severity of Illness': ['Extreme', 'Minor', 'Moderate'],
'Stay': ['0-10',
'11-20',
'21-30',
'31-40',
'41-50',
'51-60',
'61-70',
'71-80',
'81-90',
'91-100',
'More than 100 Days'],
'Type of Admission': ['Emergency', 'Trauma', 'Urgent']}
unique_list = categorical_uniques['Type of Admission']
unique_list.insert(0, unique_list.pop(unique_list.index('Urgent')))
unique_list.insert(0, unique_list.pop(unique_list.index('Trauma')))
unique_list = categorical_uniques['Severity of Illness']
unique_list.insert(0, unique_list.pop(unique_list.index('Moderate')))
unique_list.insert(0, unique_list.pop(unique_list.index('Minor')))
categorical_uniques
{'Age': ['0-10',
'11-20',
'21-30',
'31-40',
'41-50',
'51-60',
'61-70',
'71-80',
'81-90',
'91-100'],
'Severity of Illness': ['Minor', 'Moderate', 'Extreme'],
'Stay': ['0-10',
'11-20',
'21-30',
'31-40',
'41-50',
'51-60',
'61-70',
'71-80',
'81-90',
'91-100',
'More than 100 Days'],
'Type of Admission': ['Trauma', 'Urgent', 'Emergency']}
stay_mappings = {value: index for index, value in enumerate(categorical_uniques['Stay'])}
def ordinal_encode(data, uniques):
for column in uniques:
data[column] = data[column].apply(lambda x: uniques[column].index(x))
ordinal_encode(data, categorical_uniques)
data['Stay']
0 0
1 4
2 3
3 4
4 4
..
318433 1
318434 3
318435 1
318436 1
318437 0
Name: Stay, Length: 318438, dtype: int64
すべてが数値になっていることを確認
(data.dtypes == 'object').sum()
0
スケーリング
data = data.set_index('case_id')
y = data['Stay']
X = data.drop(['Stay'], axis=1)
y
case_id
1 0
2 4
3 3
4 4
5 4
..
318434 1
318435 3
318436 1
318437 1
318438 0
Name: Stay, Length: 318438, dtype: int64
scaler = sp.StandardScaler()
X = pd.DataFrame(scaler.fit_transform(X), index=X.index, columns=X.columns)
X
| Hospital_code | City_Code_Hospital | Available Extra Rooms in Hospital | Bed Grade | patientid | City_Code_Patient | Type of Admission | Severity of Illness | Visitors with Patient | Age | Admission_Deposit | TB & Chest disease | anesthesia | gynecology | radiotherapy | surgery | X | Y | Z | a | b | c | d | e | f | g | A | B | C | D | E | F | P | Q | R | S | T | U | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| case_id | ||||||||||||||||||||||||||||||||||||||
| 1 | -1.195176 | -0.571055 | -0.169177 | -0.716855 | -0.904442 | -0.053458 | 1.212557 | 1.646648 | -0.727923 | 0.461600 | 0.027835 | -0.176175 | -0.320416 | -1.902171 | 3.188572 | -0.061529 | -0.848727 | -0.790317 | 2.020115 | -0.905268 | -0.525686 | 2.435861 | -0.26155 | -0.290425 | -0.186494 | -0.116679 | -0.309922 | -0.352282 | -0.354009 | -0.440807 | -0.458683 | 1.350633 | -0.126891 | -0.707202 | 1.220175 | -0.568572 | -0.068263 | -0.005316 |
| 2 | -1.890124 | 0.073580 | -1.025217 | -0.716855 | -0.904442 | -0.053458 | -0.974973 | 1.646648 | -0.727923 | 0.461600 | 0.987556 | -0.176175 | -0.320416 | -1.902171 | 3.188572 | -0.061529 | -0.848727 | -0.790317 | 2.020115 | -0.905268 | -0.525686 | 2.435861 | -0.26155 | -0.290425 | -0.186494 | -0.116679 | -0.309922 | -0.352282 | -0.354009 | -0.440807 | -0.458683 | 1.350633 | -0.126891 | -0.707202 | -0.819554 | 1.758792 | -0.068263 | -0.005316 |
| 3 | -0.963527 | -1.215691 | -1.025217 | -0.716855 | -0.904442 | -0.053458 | -0.974973 | 1.646648 | -0.727923 | 0.461600 | -0.124910 | -0.176175 | 3.120939 | -1.902171 | -0.313620 | -0.061529 | 1.178235 | -0.790317 | -0.495021 | -0.905268 | -0.525686 | -0.410533 | -0.26155 | 3.443224 | -0.186494 | -0.116679 | -0.309922 | -0.352282 | -0.354009 | -0.440807 | 2.180153 | -0.740394 | -0.126891 | -0.707202 | -0.819554 | 1.758792 | -0.068263 | -0.005316 |
| 4 | 0.889668 | -0.893373 | -1.025217 | -0.716855 | -0.904442 | -0.053458 | -0.974973 | 1.646648 | -0.727923 | 0.461600 | 2.200319 | -0.176175 | -0.320416 | -1.902171 | 3.188572 | -0.061529 | -0.848727 | 1.265315 | -0.495021 | -0.905268 | 1.902277 | -0.410533 | -0.26155 | -0.290425 | -0.186494 | -0.116679 | -0.309922 | -0.352282 | -0.354009 | 2.268564 | -0.458683 | -0.740394 | -0.126891 | -0.707202 | 1.220175 | -0.568572 | -0.068263 | -0.005316 |
| 5 | 0.889668 | -0.893373 | -1.025217 | -0.716855 | -0.904442 | -0.053458 | -0.974973 | 1.646648 | -0.727923 | 0.461600 | 0.623175 | -0.176175 | -0.320416 | -1.902171 | 3.188572 | -0.061529 | -0.848727 | 1.265315 | -0.495021 | -0.905268 | 1.902277 | -0.410533 | -0.26155 | -0.290425 | -0.186494 | -0.116679 | -0.309922 | -0.352282 | -0.354009 | 2.268564 | -0.458683 | -0.740394 | -0.126891 | -0.707202 | -0.819554 | 1.758792 | -0.068263 | -0.005316 |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 318434 | -1.426825 | 0.395897 | -0.169177 | 1.574123 | 0.546379 | 3.342582 | 1.212557 | 0.138090 | -0.161049 | -0.067622 | -0.677923 | -0.176175 | -0.320416 | -1.902171 | 3.188572 | -0.061529 | 1.178235 | -0.790317 | -0.495021 | 1.104645 | -0.525686 | -0.410533 | -0.26155 | -0.290425 | -0.186494 | -0.116679 | -0.309922 | -0.352282 | -0.354009 | -0.440807 | -0.458683 | 1.350633 | -0.126891 | 1.414024 | -0.819554 | -0.568572 | -0.068263 | -0.005316 |
| 318435 | 0.658018 | -1.215691 | -1.025217 | 1.574123 | -1.722559 | 0.158795 | 0.118792 | 0.138090 | 0.405826 | 2.049268 | 1.673071 | -0.176175 | 3.120939 | -1.902171 | -0.313620 | -0.061529 | 1.178235 | -0.790317 | -0.495021 | 1.104645 | -0.525686 | -0.410533 | -0.26155 | -0.290425 | -0.186494 | -0.116679 | -0.309922 | -0.352282 | -0.354009 | -0.440807 | 2.180153 | -0.740394 | -0.126891 | 1.414024 | -0.819554 | -0.568572 | -0.068263 | -0.005316 |
| 318436 | -1.311001 | -0.248738 | -0.169177 | 1.574123 | 1.566288 | 0.583300 | 1.212557 | -1.370469 | -0.161049 | 1.520045 | -0.594189 | -0.176175 | -0.320416 | 0.525715 | -0.313620 | -0.061529 | 1.178235 | -0.790317 | -0.495021 | 1.104645 | -0.525686 | -0.410533 | -0.26155 | -0.290425 | -0.186494 | -0.116679 | -0.309922 | -0.352282 | -0.354009 | -0.440807 | -0.458683 | 1.350633 | -0.126891 | -0.707202 | 1.220175 | -0.568572 | -0.068263 | -0.005316 |
| 318437 | -0.847702 | -0.893373 | -0.169177 | 0.428634 | 0.667022 | 0.158795 | -0.974973 | -1.370469 | 0.972701 | -1.655290 | -1.030342 | -0.176175 | 3.120939 | -1.902171 | -0.313620 | -0.061529 | -0.848727 | 1.265315 | -0.495021 | -0.905268 | 1.902277 | -0.410533 | -0.26155 | -0.290425 | -0.186494 | -0.116679 | -0.309922 | -0.352282 | -0.354009 | 2.268564 | -0.458683 | -0.740394 | -0.126891 | 1.414024 | -0.819554 | -0.568572 | -0.068263 | -0.005316 |
| 318438 | 0.078895 | 0.718215 | 1.542903 | -0.716855 | -1.161314 | 0.158795 | 1.212557 | -1.370469 | -0.727923 | -1.655290 | -0.118469 | -0.176175 | -0.320416 | 0.525715 | -0.313620 | -0.061529 | -0.848727 | 1.265315 | -0.495021 | 1.104645 | -0.525686 | -0.410533 | -0.26155 | -0.290425 | -0.186494 | -0.116679 | -0.309922 | -0.352282 | 2.824791 | -0.440807 | -0.458683 | -0.740394 | -0.126891 | 1.414024 | -0.819554 | -0.568572 | -0.068263 | -0.005316 |
318438 rows × 38 columns
トレーニング
X_train, X_test, y_train, y_test = train_test_split(X, y, train_size=0.8)
log_model = LogisticRegression()
log_model.fit(X_train, y_train)
/usr/local/lib/python3.6/dist-packages/sklearn/linear_model/_logistic.py:940: ConvergenceWarning: lbfgs failed to converge (status=1):
STOP: TOTAL NO. of ITERATIONS REACHED LIMIT.
Increase the number of iterations (max_iter) or scale the data as shown in:
https://scikit-learn.org/stable/modules/preprocessing.html
Please also refer to the documentation for alternative solver options:
https://scikit-learn.org/stable/modules/linear_model.html#logistic-regression
extra_warning_msg=_LOGISTIC_SOLVER_CONVERGENCE_MSG)
LogisticRegression(C=1.0, class_weight=None, dual=False, fit_intercept=True,
intercept_scaling=1, l1_ratio=None, max_iter=100,
multi_class='auto', n_jobs=None, penalty='l2',
random_state=None, solver='lbfgs', tol=0.0001, verbose=0,
warm_start=False)
nn_model = tf.keras.Sequential([
tf.keras.layers.Dense(16, activation='relu', input_shape=(38, )),
tf.keras.layers.Dense(16, activation='relu'),
tf.keras.layers.Dense(11, activation='softmax'),
])
nn_model.summary()
Model: "sequential_1"
_________________________________________________________________
Layer (type) Output Shape Param #
=================================================================
dense_3 (Dense) (None, 16) 624
_________________________________________________________________
dense_4 (Dense) (None, 16) 272
_________________________________________________________________
dense_5 (Dense) (None, 11) 187
=================================================================
Total params: 1,083
Trainable params: 1,083
Non-trainable params: 0
_________________________________________________________________
nn_model.compile(
optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'],
)
batch_size = 32
epochs = 10
history = nn_model.fit(
X_train,
y_train,
validation_split=0.2,
batch_size=batch_size,
epochs=epochs,
)
Epoch 1/10
6369/6369 [==============================] - 8s 1ms/step - loss: 1.6387 - accuracy: 0.3739 - val_loss: 1.5888 - val_accuracy: 0.3951
Epoch 2/10
6369/6369 [==============================] - 8s 1ms/step - loss: 1.5738 - accuracy: 0.3990 - val_loss: 1.5667 - val_accuracy: 0.4029
Epoch 3/10
6369/6369 [==============================] - 8s 1ms/step - loss: 1.5593 - accuracy: 0.4034 - val_loss: 1.5563 - val_accuracy: 0.4081
Epoch 4/10
6369/6369 [==============================] - 8s 1ms/step - loss: 1.5528 - accuracy: 0.4057 - val_loss: 1.5530 - val_accuracy: 0.4080
Epoch 5/10
6369/6369 [==============================] - 8s 1ms/step - loss: 1.5488 - accuracy: 0.4075 - val_loss: 1.5494 - val_accuracy: 0.4100
Epoch 6/10
6369/6369 [==============================] - 8s 1ms/step - loss: 1.5462 - accuracy: 0.4083 - val_loss: 1.5463 - val_accuracy: 0.4104
Epoch 7/10
6369/6369 [==============================] - 8s 1ms/step - loss: 1.5445 - accuracy: 0.4084 - val_loss: 1.5462 - val_accuracy: 0.4093
Epoch 8/10
6369/6369 [==============================] - 8s 1ms/step - loss: 1.5427 - accuracy: 0.4096 - val_loss: 1.5454 - val_accuracy: 0.4086
Epoch 9/10
6369/6369 [==============================] - 8s 1ms/step - loss: 1.5416 - accuracy: 0.4102 - val_loss: 1.5452 - val_accuracy: 0.4117
Epoch 10/10
6369/6369 [==============================] - 8s 1ms/step - loss: 1.5404 - accuracy: 0.4109 - val_loss: 1.5431 - val_accuracy: 0.4120
print(f'Logistic Regression Acc: {log_model.score(X_test, y_test)}')
print(f' Neural Network Acc: {nn_model.evaluate(X_test, y_test, verbose=0)[1]}')
Logistic Regression Acc: 0.38908428589373195
Neural Network Acc: 0.41164740920066833
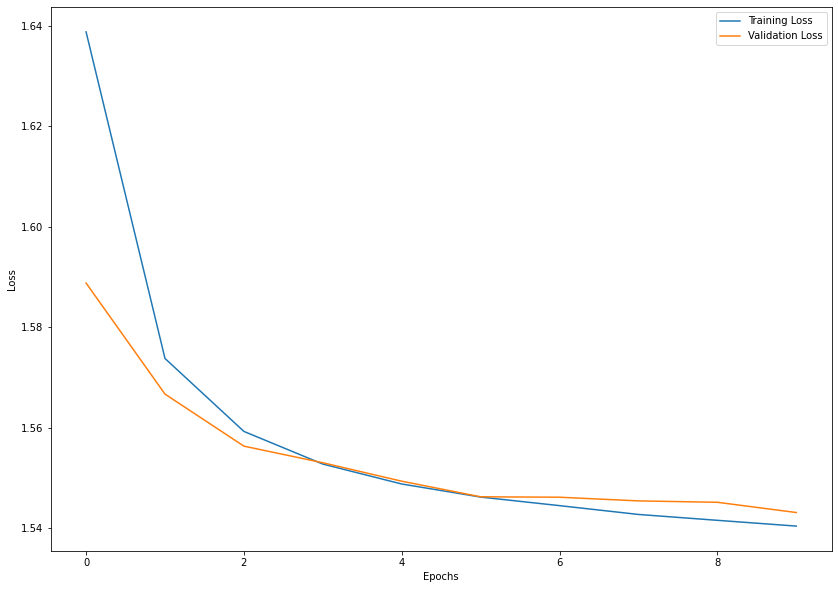
plt.figure(figsize=(14, 10))
plt.plot(range(epochs), history.history['loss'], label='Training Loss')
plt.plot(range(epochs), history.history['val_loss'], label='Validation Loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend(loc='upper right')
plt.show()
np.argmin(history.history['val_loss']) + 1
10
今回はsklearnのロジスティック回帰とtensorflowの両方でモデルを構築しました。
比較するとtensorflowのほうが若干良い精度がでていました。
NNいいですね。